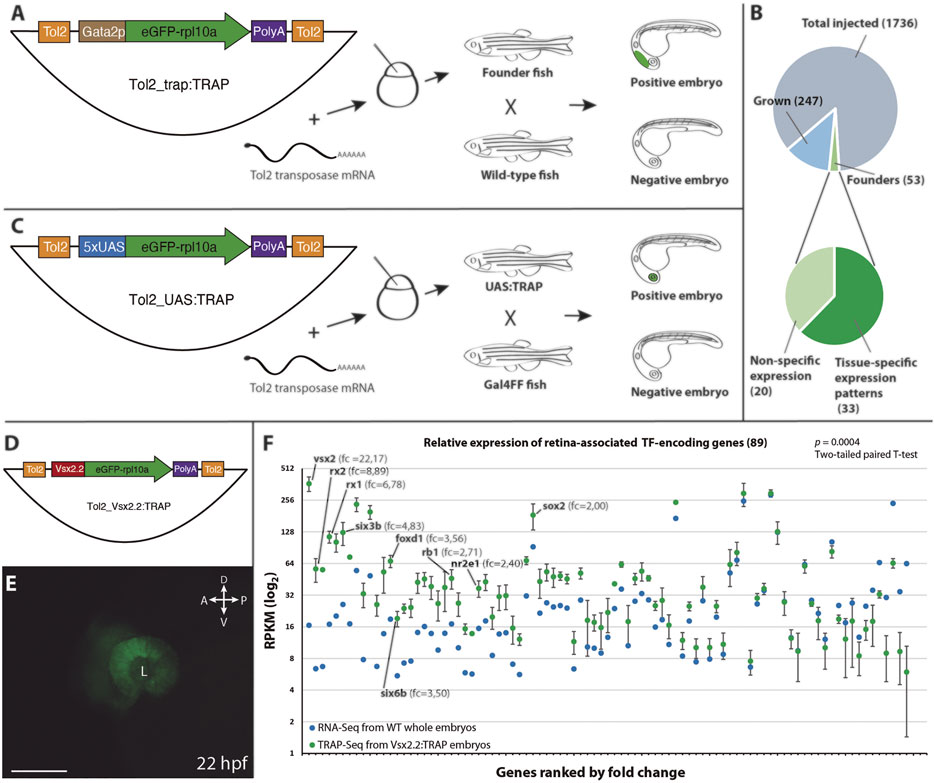
TrapSeq: An RNA Sequencing-Based Pipeline for the Identification of Gene- Trap Insertions in Mammalian Cells - ScienceDirect

Translating Ribosome Affinity Purification (TRAP) to Investigate Arabidopsis thaliana Root Development at a Cell Type-Specific Scale | Protocol (Translated to German)

Monitoring Cell-Type–Specific Gene Expression Using Ribosome Profiling In Vivo During Cardiac Hemodynamic Stress | Circulation Research

TRAP-SEQ of Eukaryotic Translatomes Applied to the Detection of Polysome-Associated Long Noncoding RNAs | SpringerLink

TRAP-SEQ – Translating Ribosome Affinity Purification (TRAP) Followed by RNA Sequencing Technology | RNA-Seq Blog
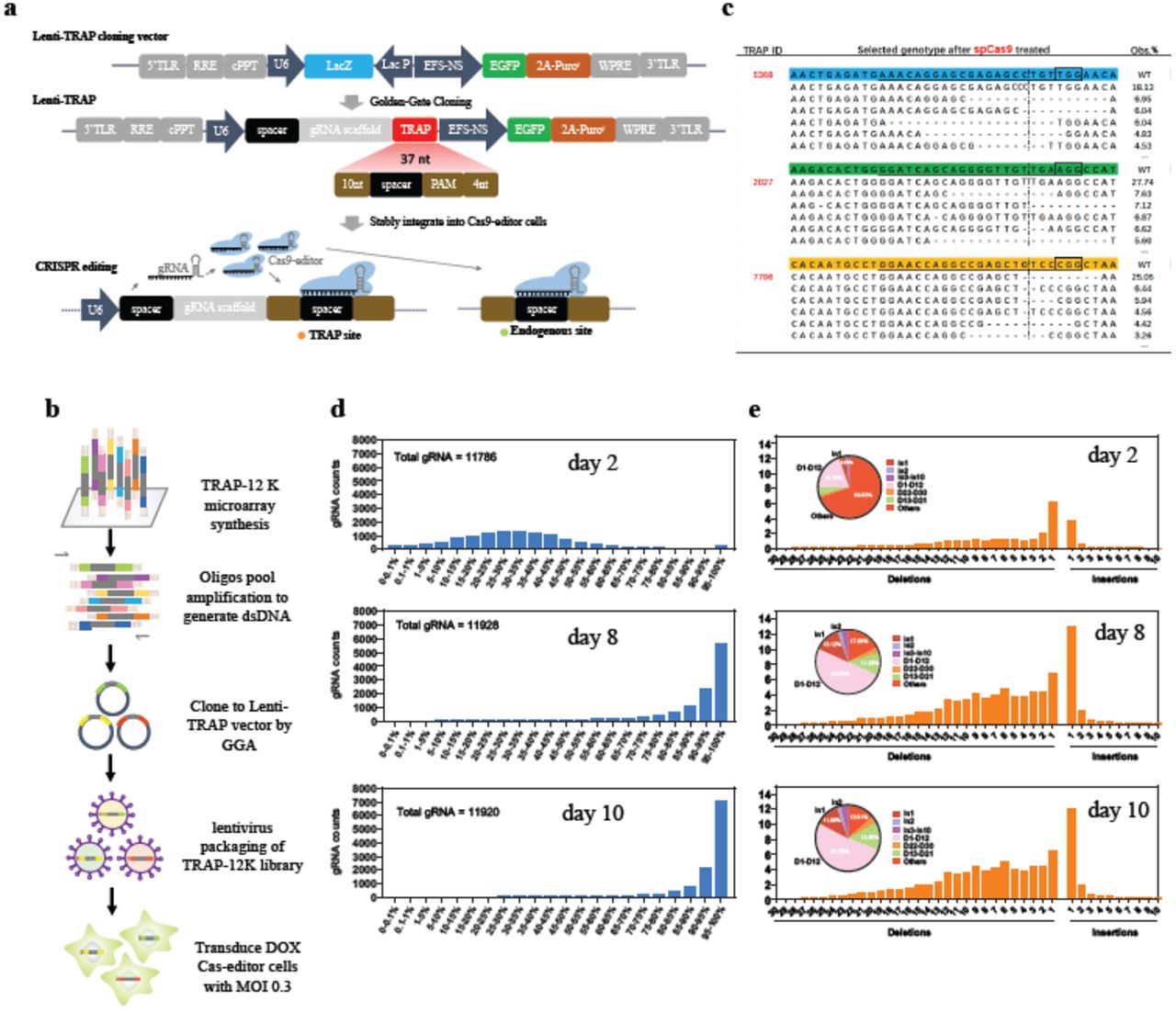
Massively parallel quantification of CRISPR editing in cells by TRAP-seq enables better design of Cas9, ABE, CBE gRNAs of high efficiency and accuracy | bioRxiv

TRAP-Seq Identifies Differentially Translating mRNAs in Fmr1 –/y CA1... | Download Scientific Diagram
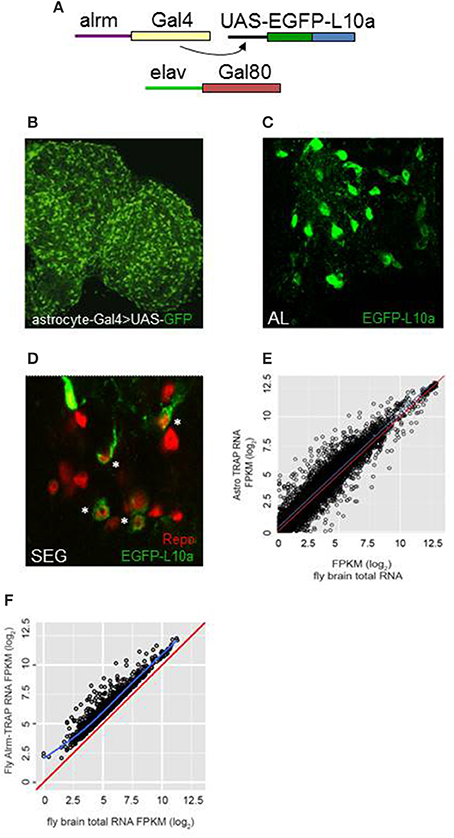
Frontiers | TRAP-seq Profiling and RNAi-Based Genetic Screens Identify Conserved Glial Genes Required for Adult Drosophila Behavior

Cell type–specific mRNA purification by translating ribosome affinity purification (TRAP) | Nature Protocols

Interrogating translational efficiency and lineage-specific transcriptomes using ribosome affinity purification | PNAS
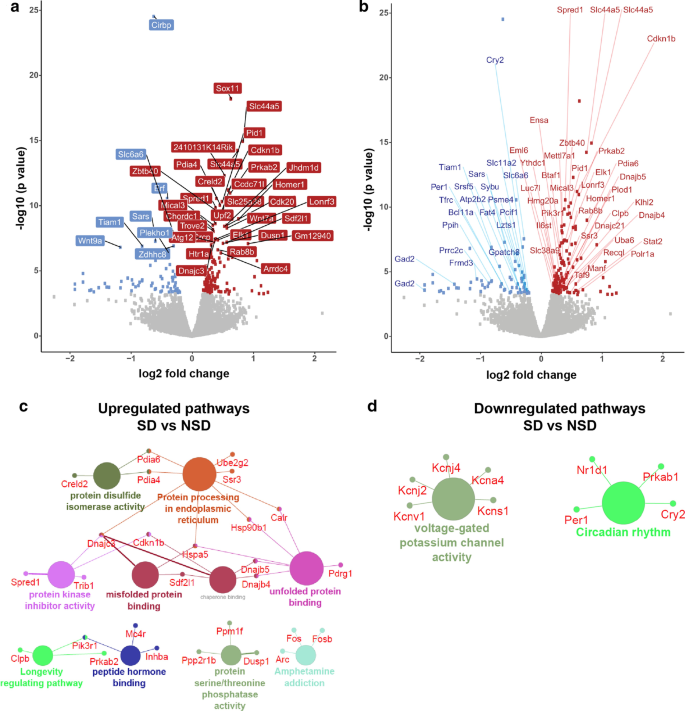
Translational changes induced by acute sleep deprivation uncovered by TRAP- Seq | Molecular Brain | Full Text

Leprechaun Trap - Story Sequencing! - Mrs. B's Beehive | Story sequencing, Sequencing activities kindergarten, Writing instruction
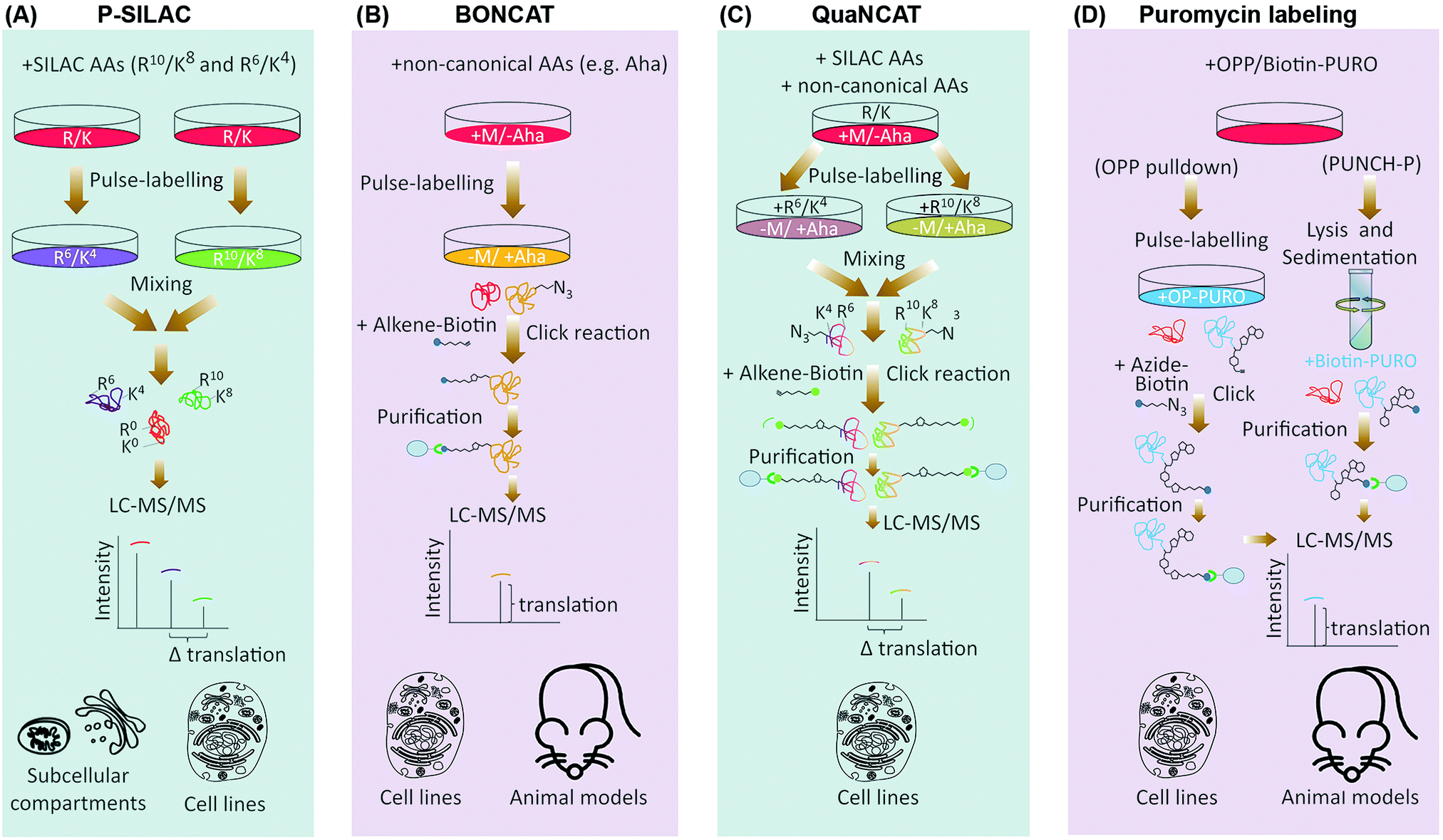
Methods for monitoring and measurement of protein translation in time and space - Molecular BioSystems (RSC Publishing) DOI:10.1039/C7MB00476A
![PDF] Evaluation of TRAP-sequencing technology with a versatile conditional mouse model | Semantic Scholar PDF] Evaluation of TRAP-sequencing technology with a versatile conditional mouse model | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/820026a2dd41a5bc8931b4d0a03ae6b567899463/7-Figure4-1.png)
PDF] Evaluation of TRAP-sequencing technology with a versatile conditional mouse model | Semantic Scholar