DNA constructs for ribosome display. (A) Construct for Escherichia coli... | Download Scientific Diagram

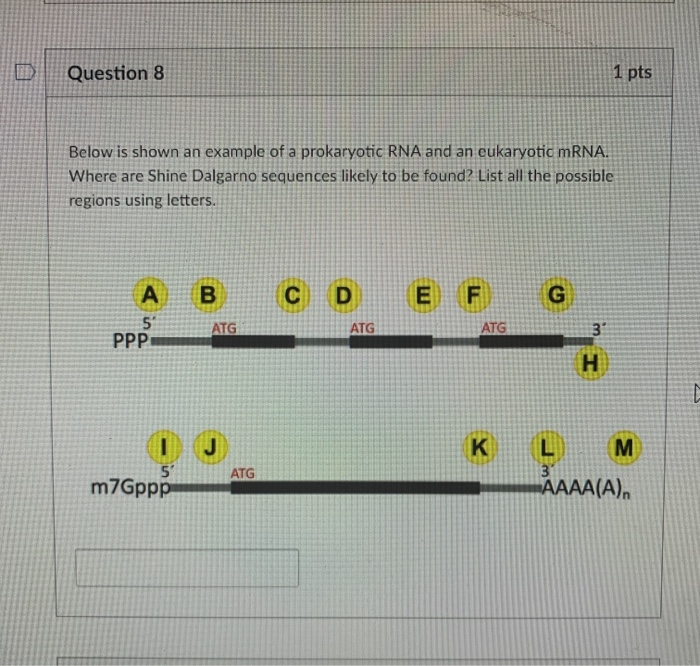
Silent mutations in the Shine-Dalgarno-like sequence and the predicted... | Download Scientific Diagram
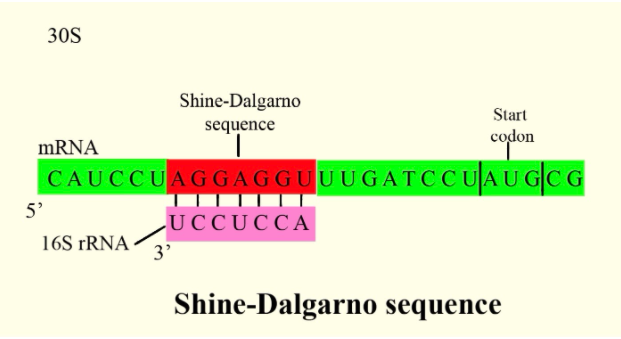
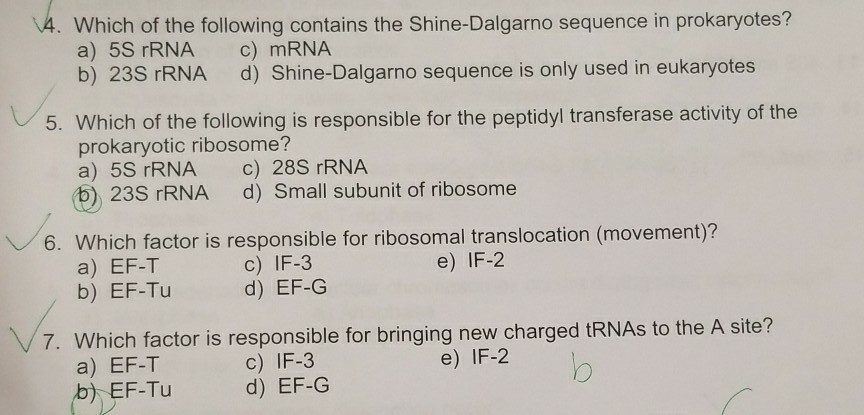
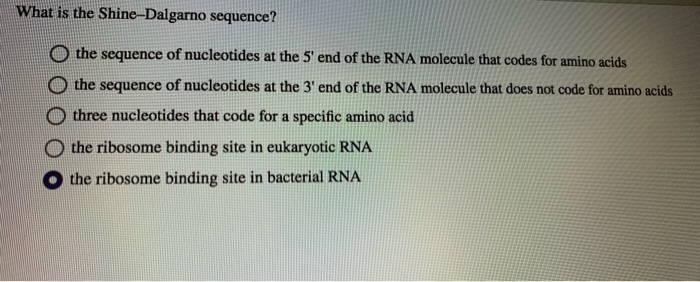
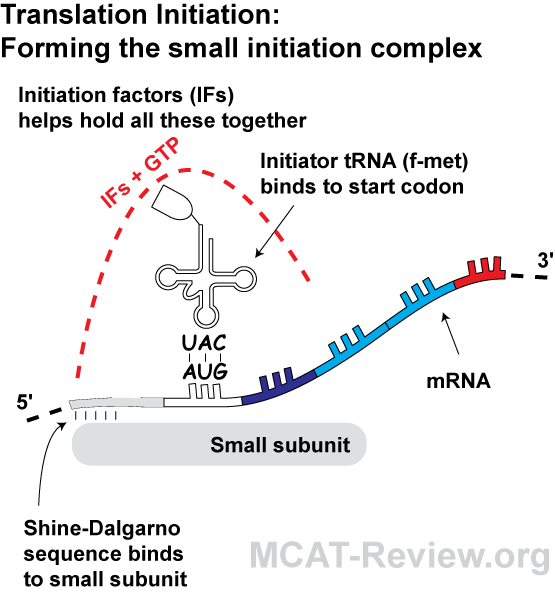
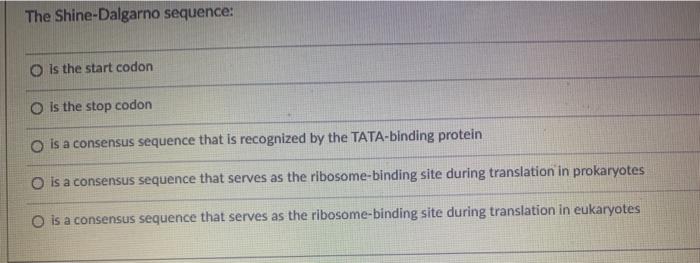
In prokaryotes, the ribosomes binding site on mRNA is called(a)Hogness sequence(b)Shine- Dalgarno sequence(c)Prinbow sequence(d)TATA- box

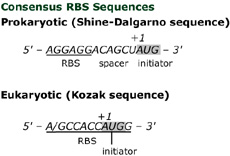
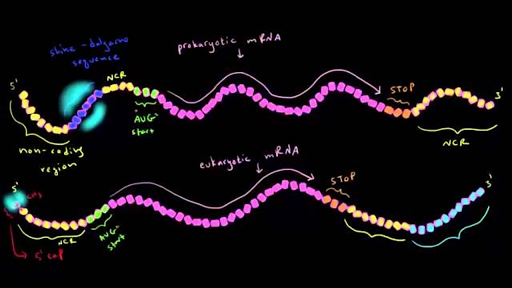
What is the Difference Between Shine Dalgarno and Kozak Sequence | Compare the Difference Between Similar Terms

The possible dual impacts of Shine-Dalgarno (SD) sequences on protein... | Download Scientific Diagram

SOLVED: Which one of the following is found in prokaryotes but not eukaryotes? 5' cap of mRNA small subunit rRNAs altemnatively spliced RNA transcripts polyadenylation Shine-Dalgarno sequence

Extending the Spacing between the Shine–Dalgarno Sequence and P-Site Codon Reduces the Rate of mRNA Translocation - ScienceDirect