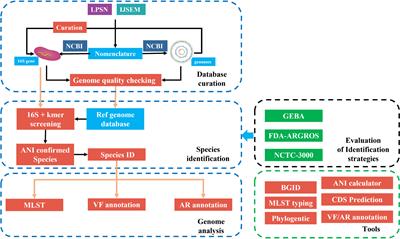
Transcriptional adaptation of olfactory sensory neurons to GPCR identity and activity | Nature Communications
Using Sequence Similarity Networks for Visualization of Relationships Across Diverse Protein Superfamilies | PLOS ONE

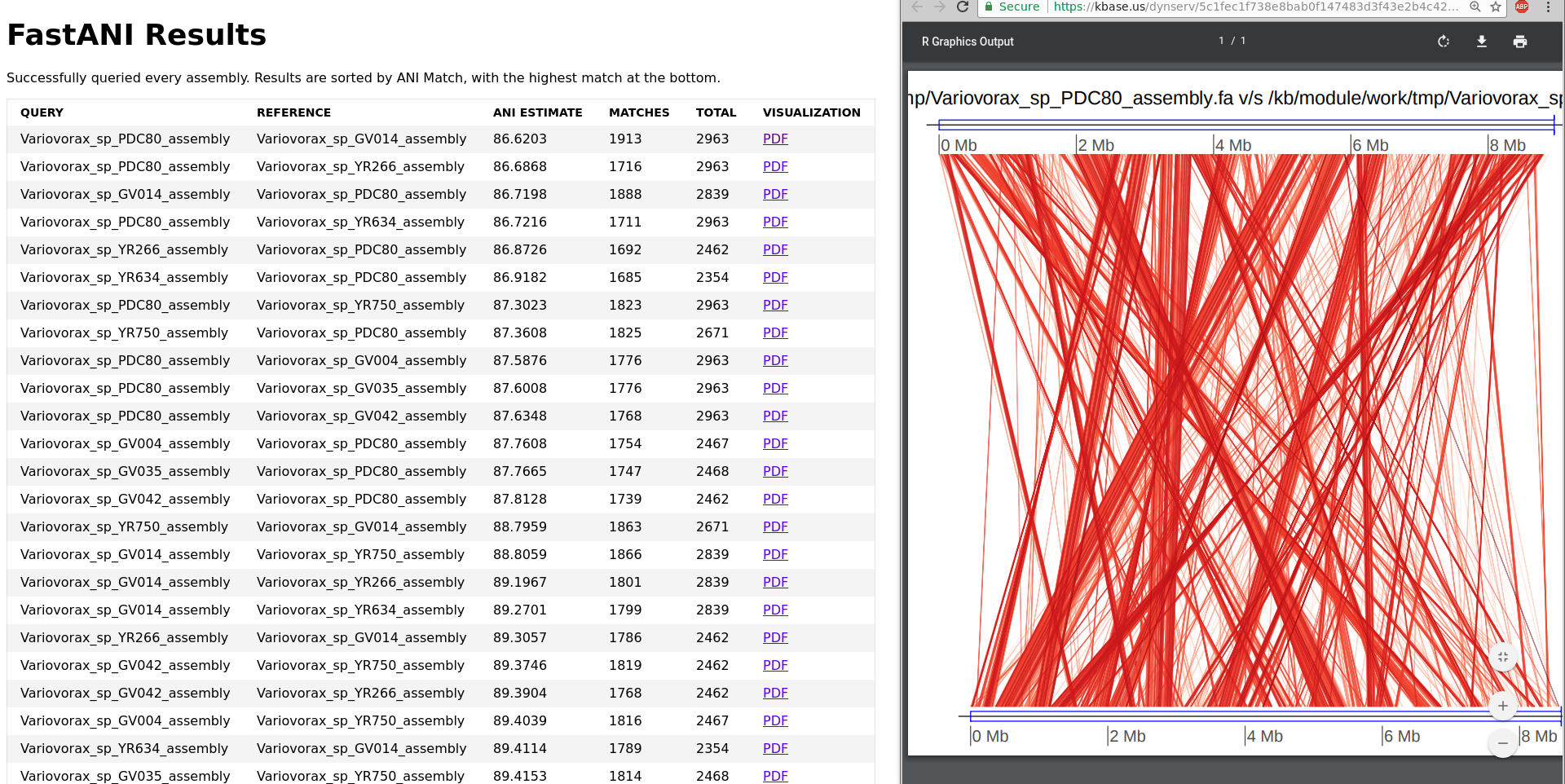
OrthoANI: An improved algorithm and software for calculating average nucleotide identity. | Semantic Scholar

How to find protein identity and similarities | Sequence homology | SIAS | Gene identity similarity - YouTube
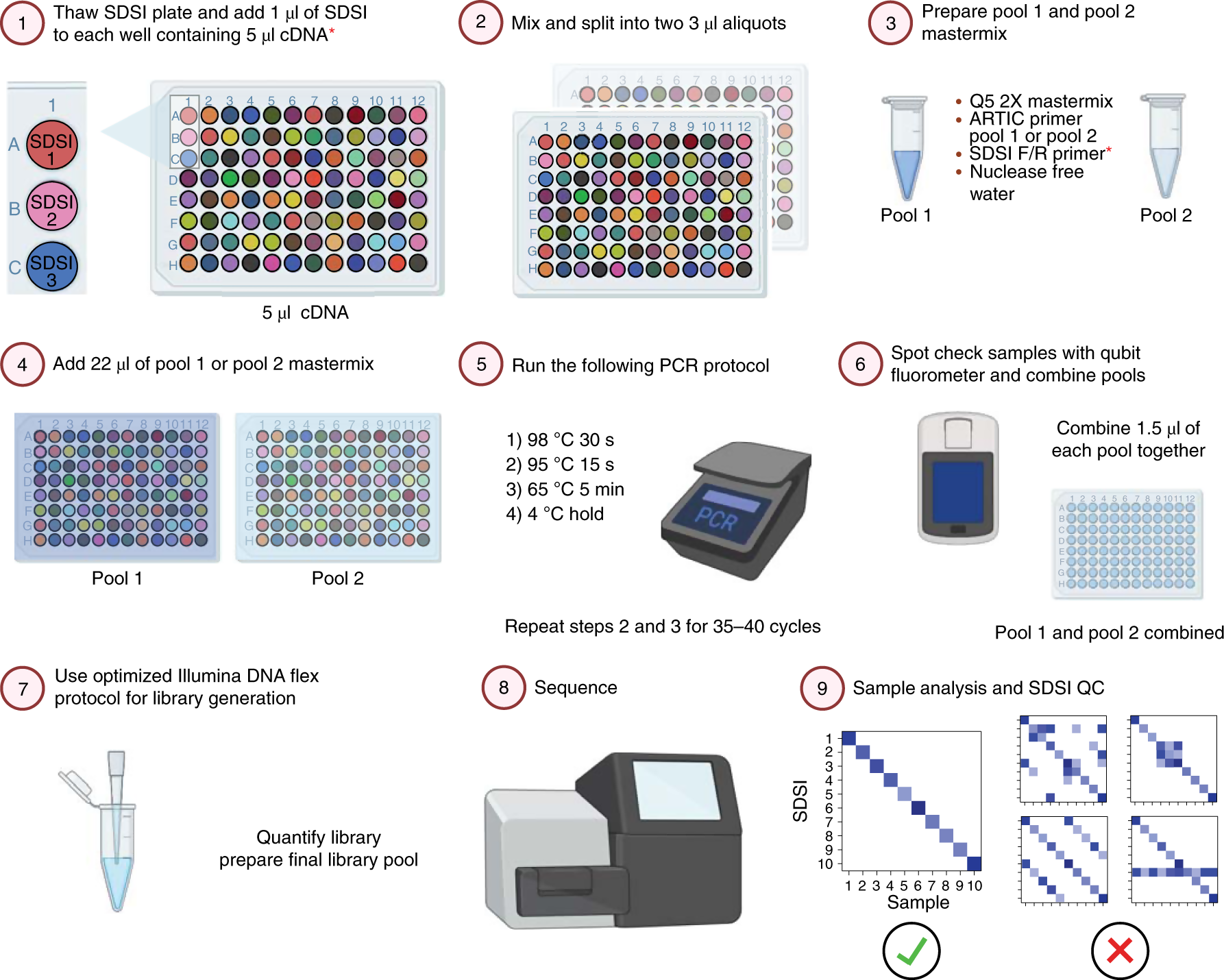
Synthetic DNA spike-ins (SDSIs) enable sample tracking and detection of inter-sample contamination in SARS-CoV-2 sequencing workflows | Nature Microbiology
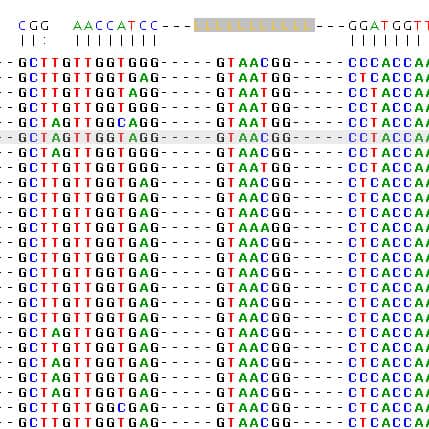
How to calculate 16S rRNA sequence similarity values for bacterial taxonomy: Why BLAST should be avoided – EzBioCloud Help center

Center for Lightweight-Production-Technology - The Dispersion Calculator: An open source software for calculating dispersion curves and mode shapes of guided waves

OrthoANI: An improved algorithm and software for calculating average nucleotide identity | Microbiology Society

How to find protein identity and similarities | Sequence homology | SIAS | Gene identity similarity - YouTube