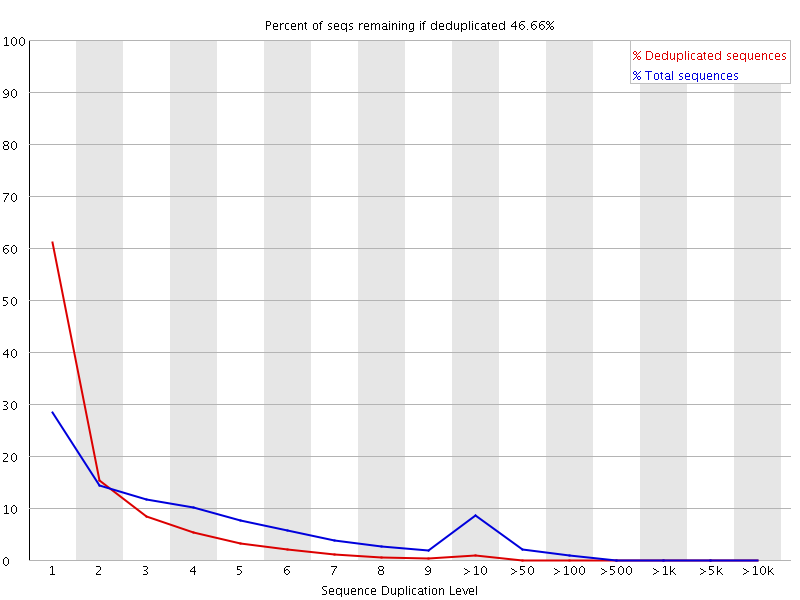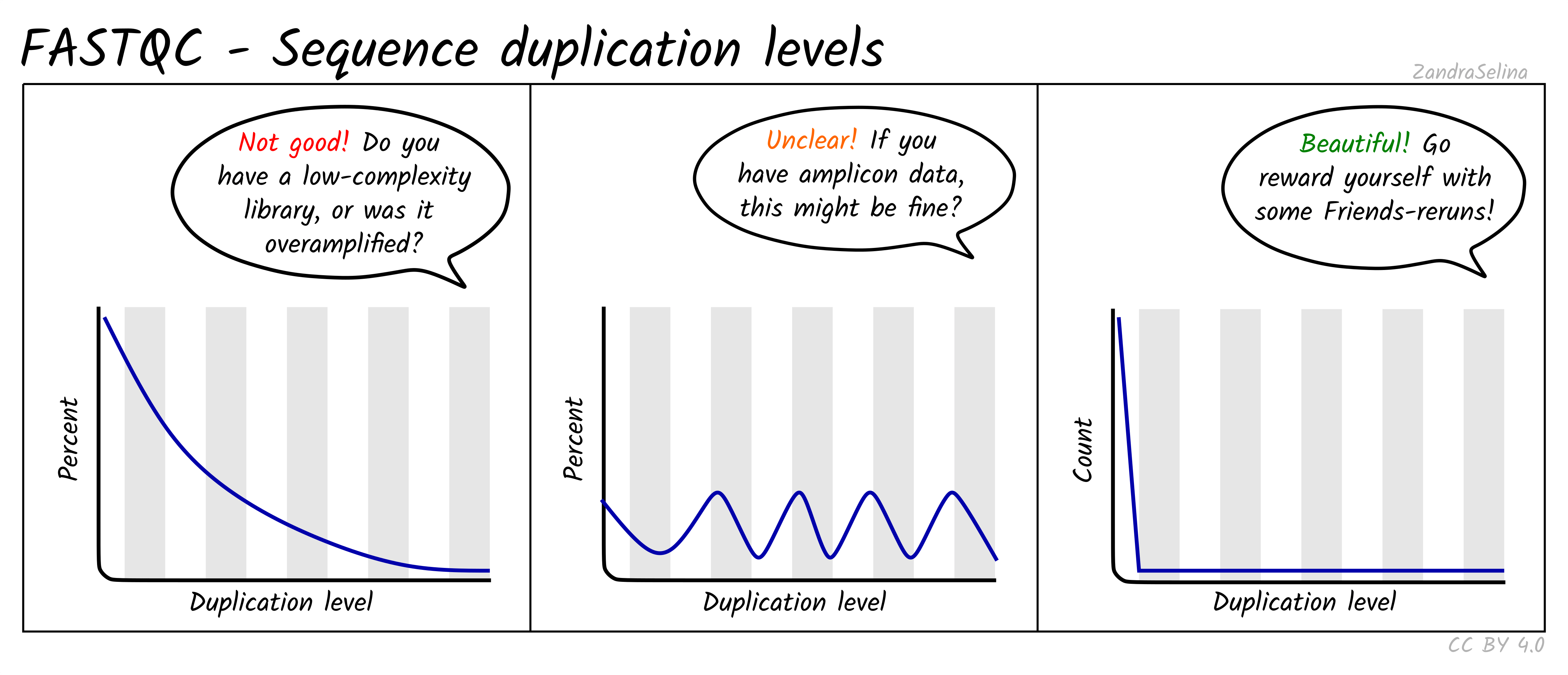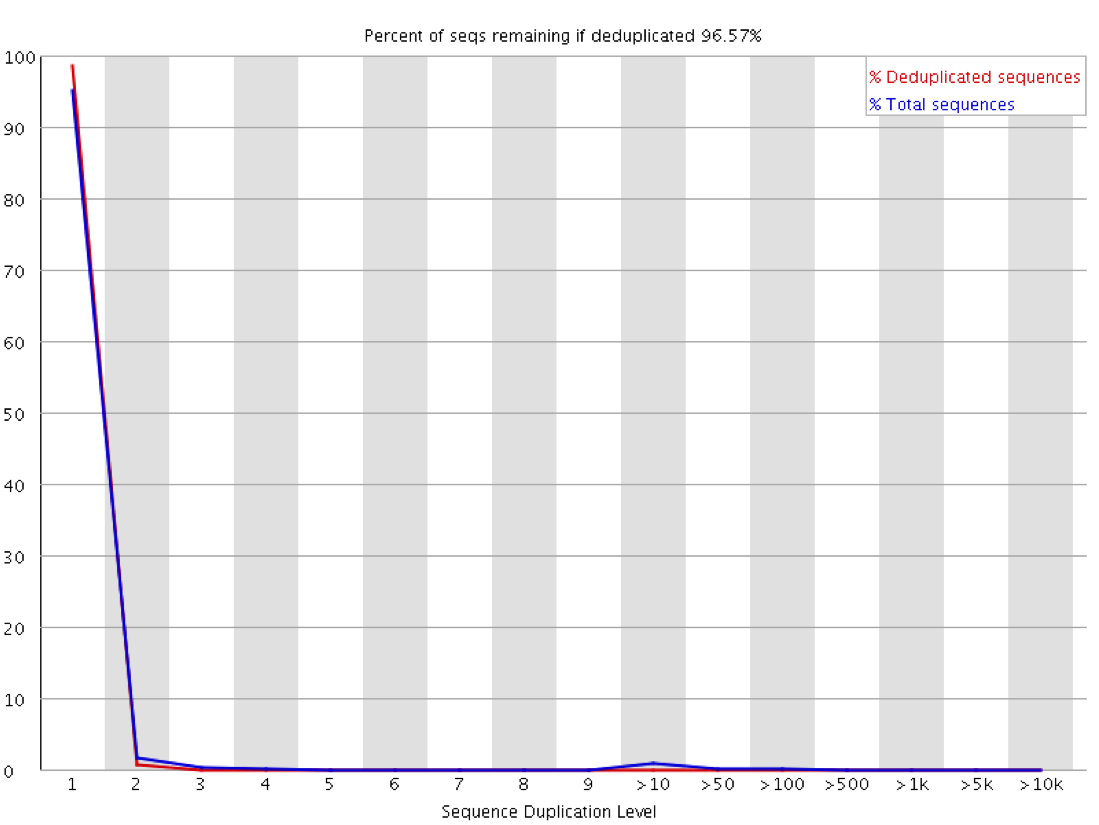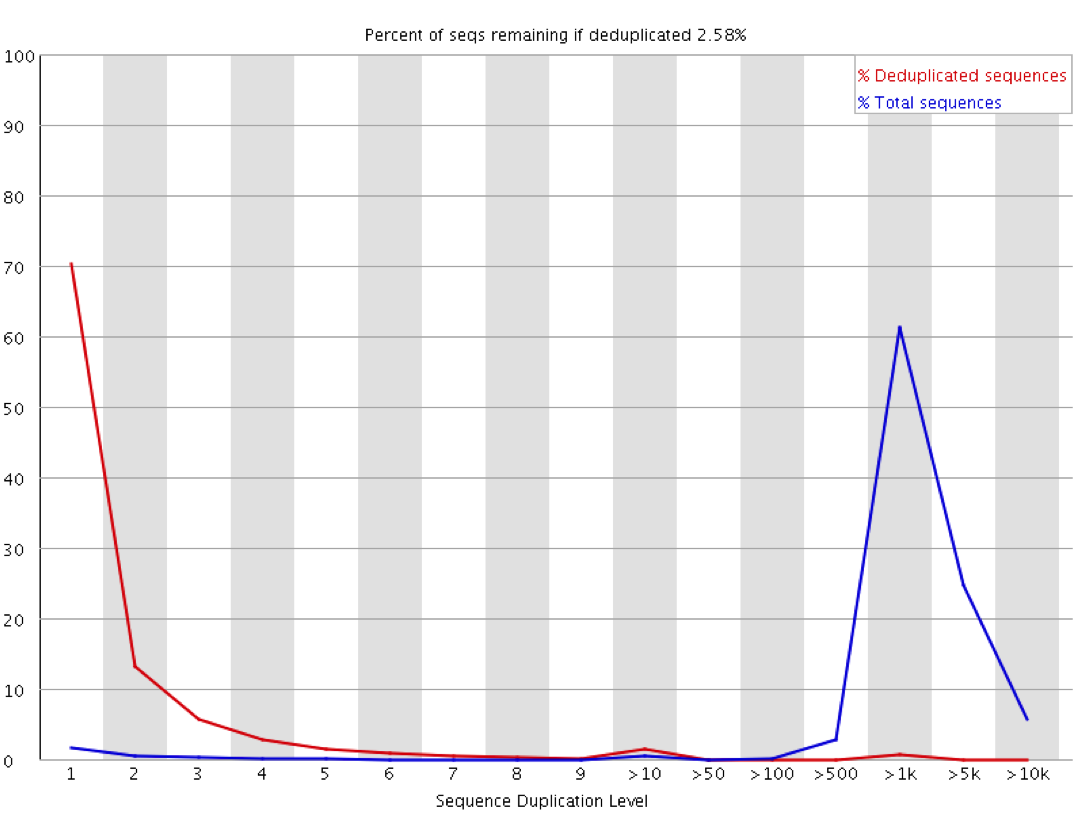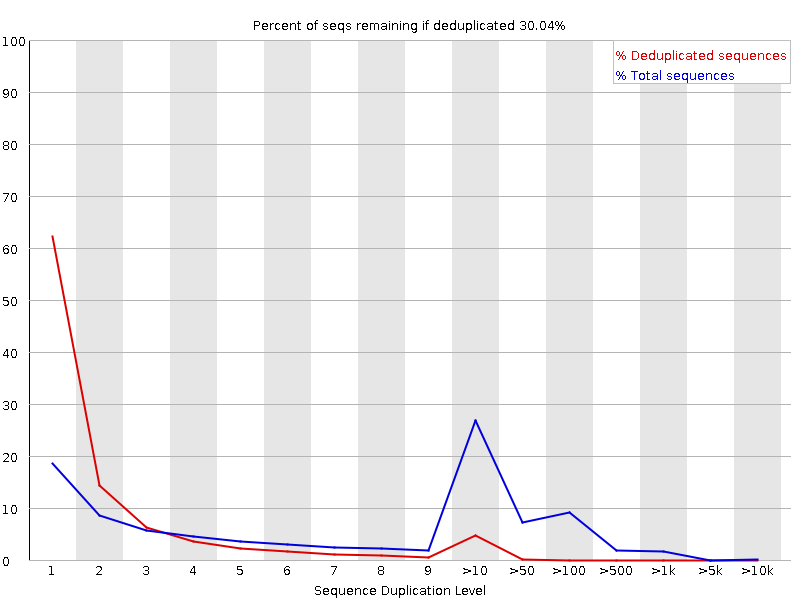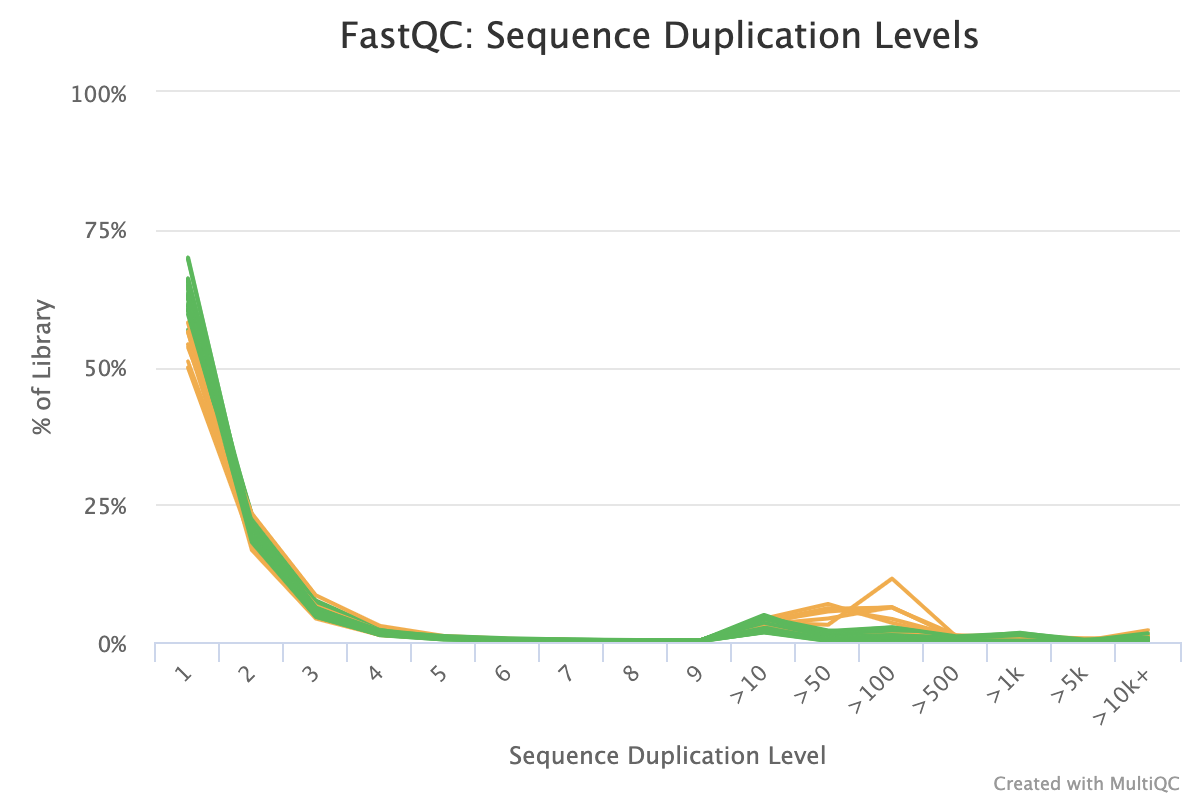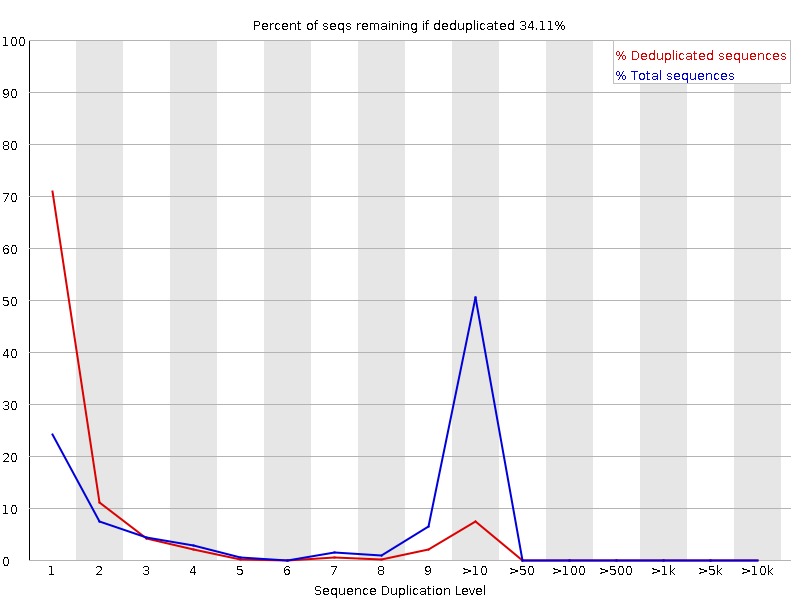
Why does FASTQC show unexpectedly high sequence duplication levels (PCR-duplicates)? | DNA Technologies Core
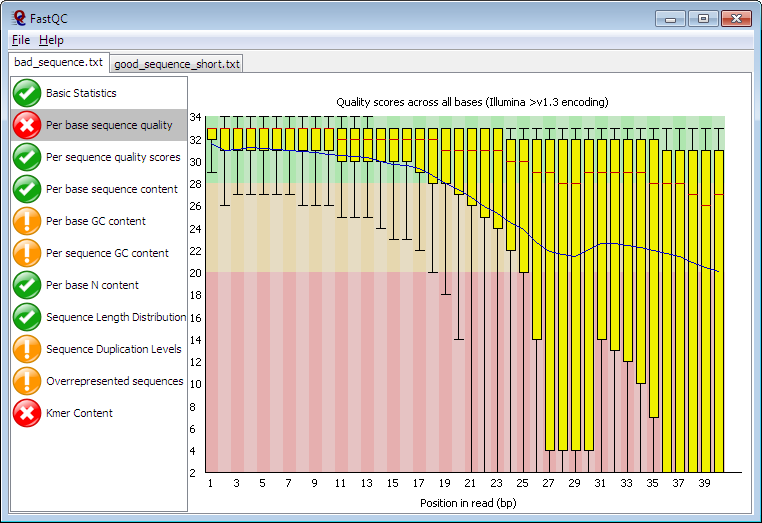
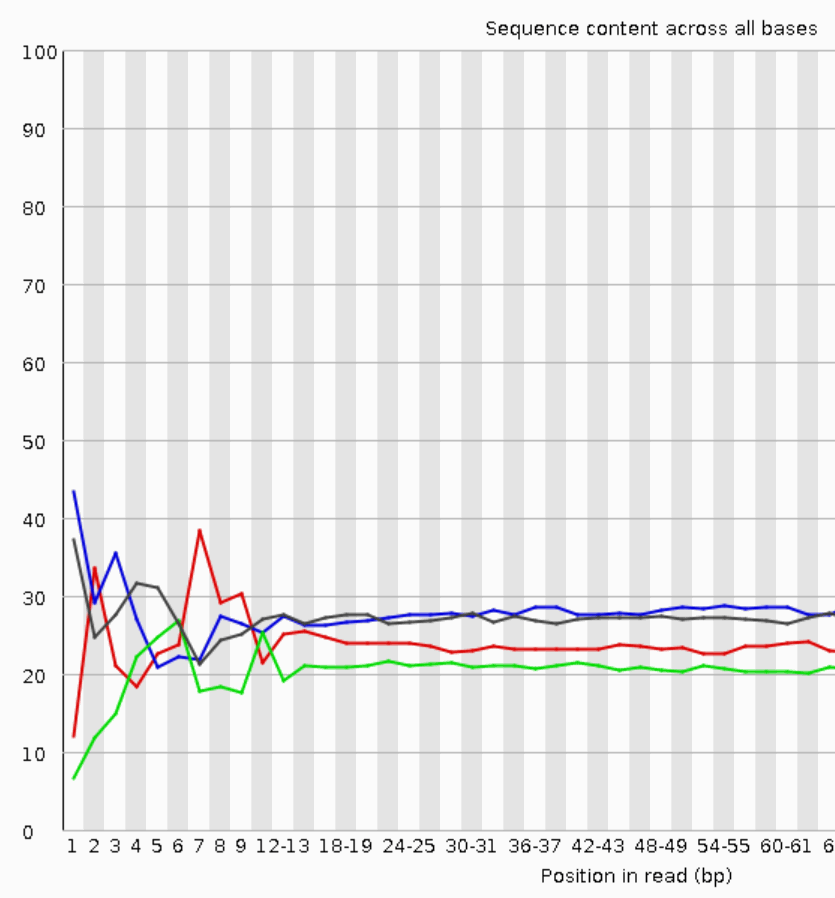
FastQC for RNA-Seq-Fails for per base seq content, gc content, duplication levels, adapter content and kmer content? | ResearchGate

FastQC: Sequence Duplication Levels interactive dialog indexing error · Issue #1092 · ewels/MultiQC · GitHub