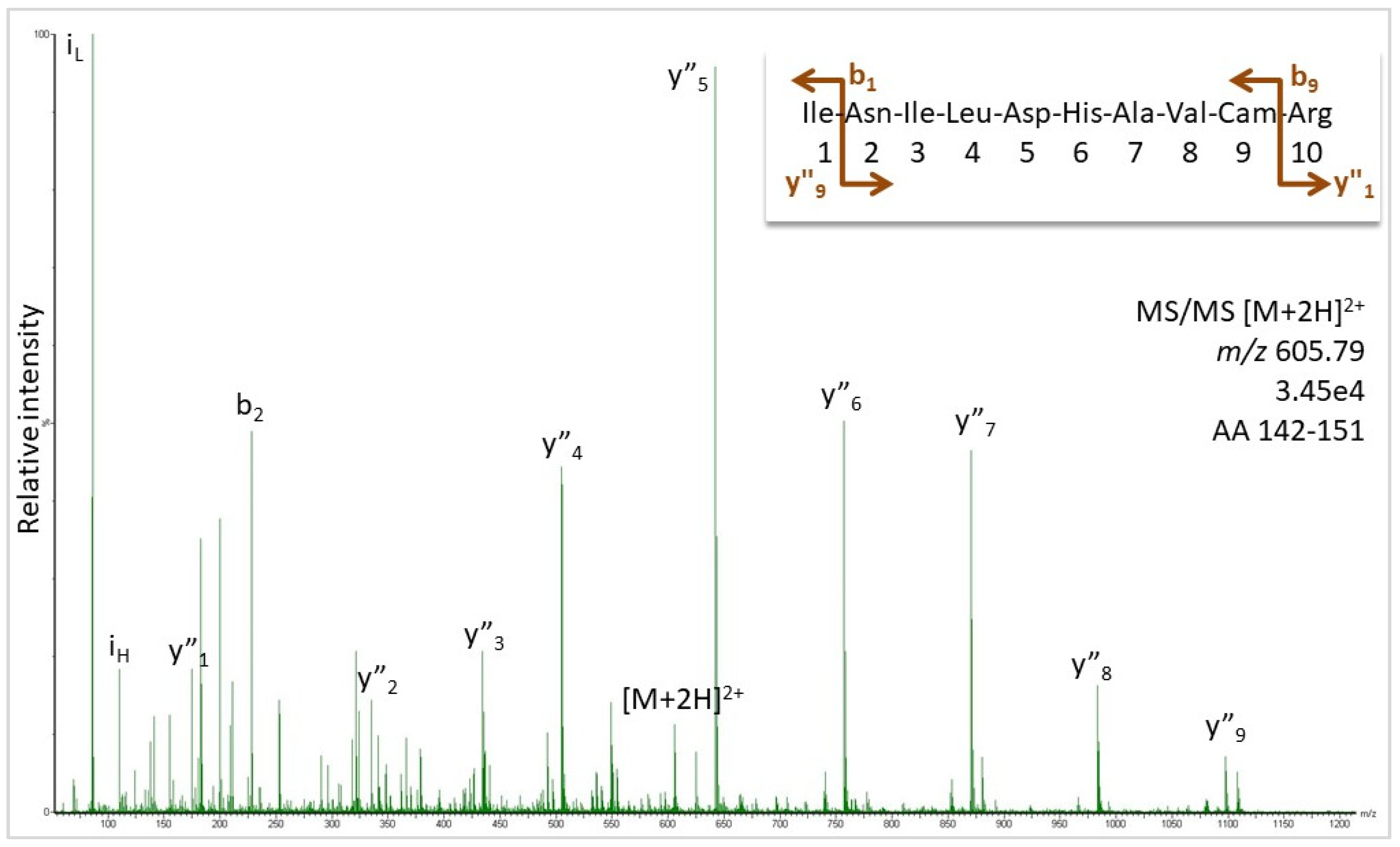
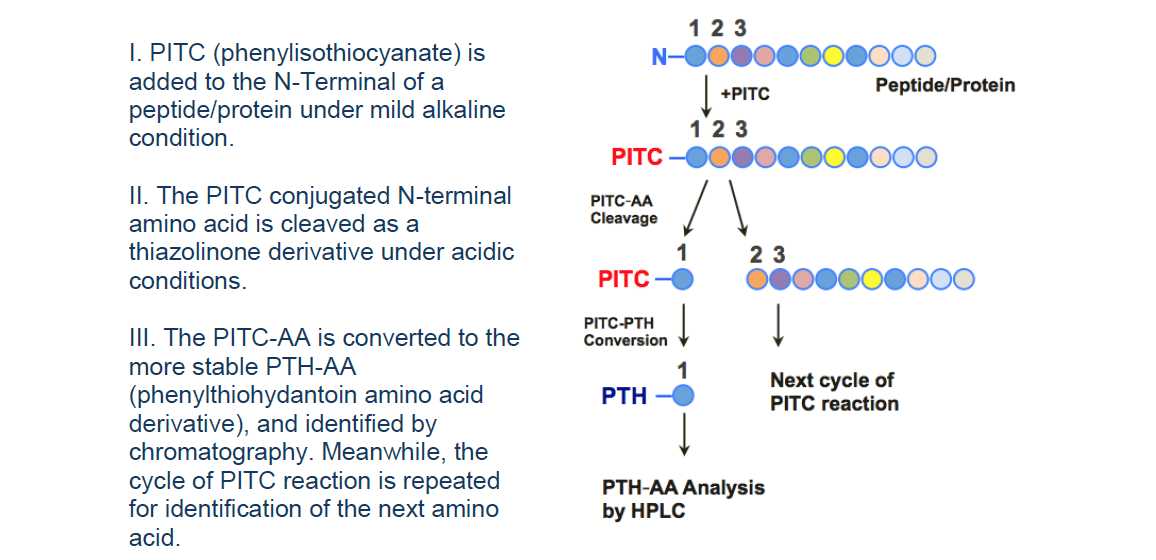
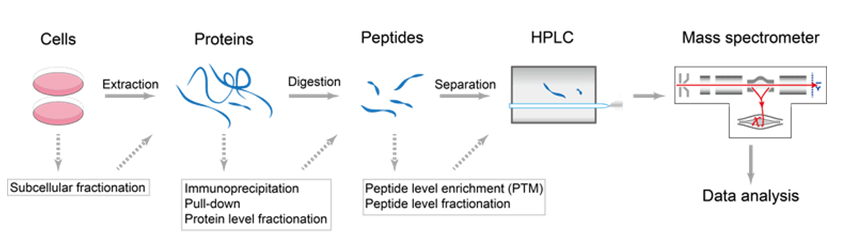
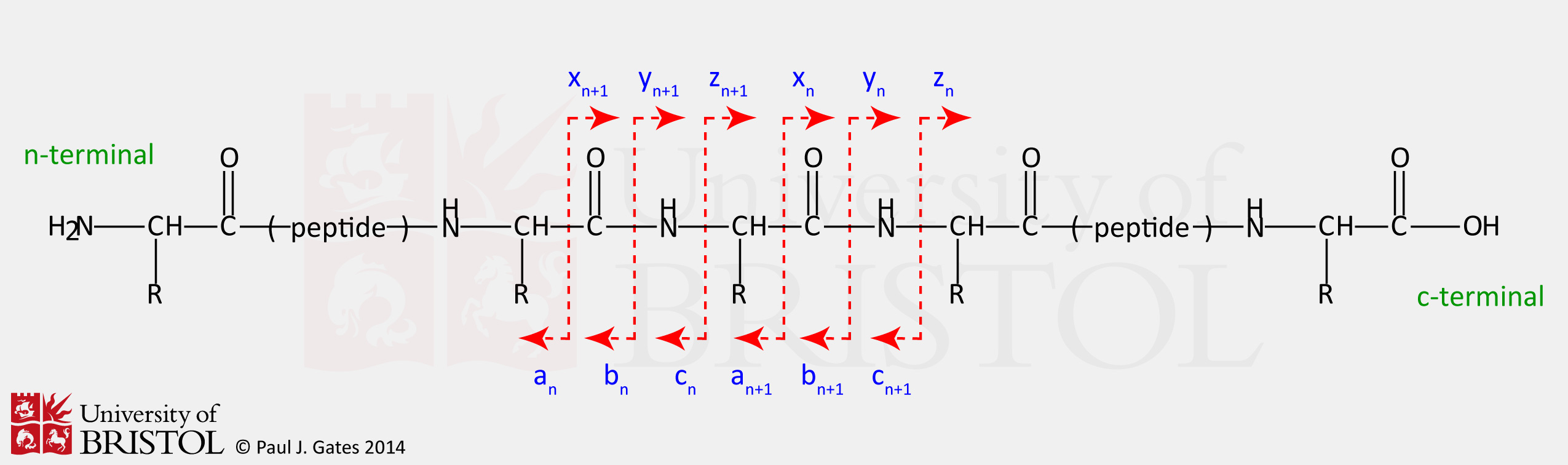
Identification and Sequencing of N-Terminal Peptides in Proteins by LC-Fluorescence-MS/MS: An Approach to Replacement of the Edman Degradation | Analytical Chemistry

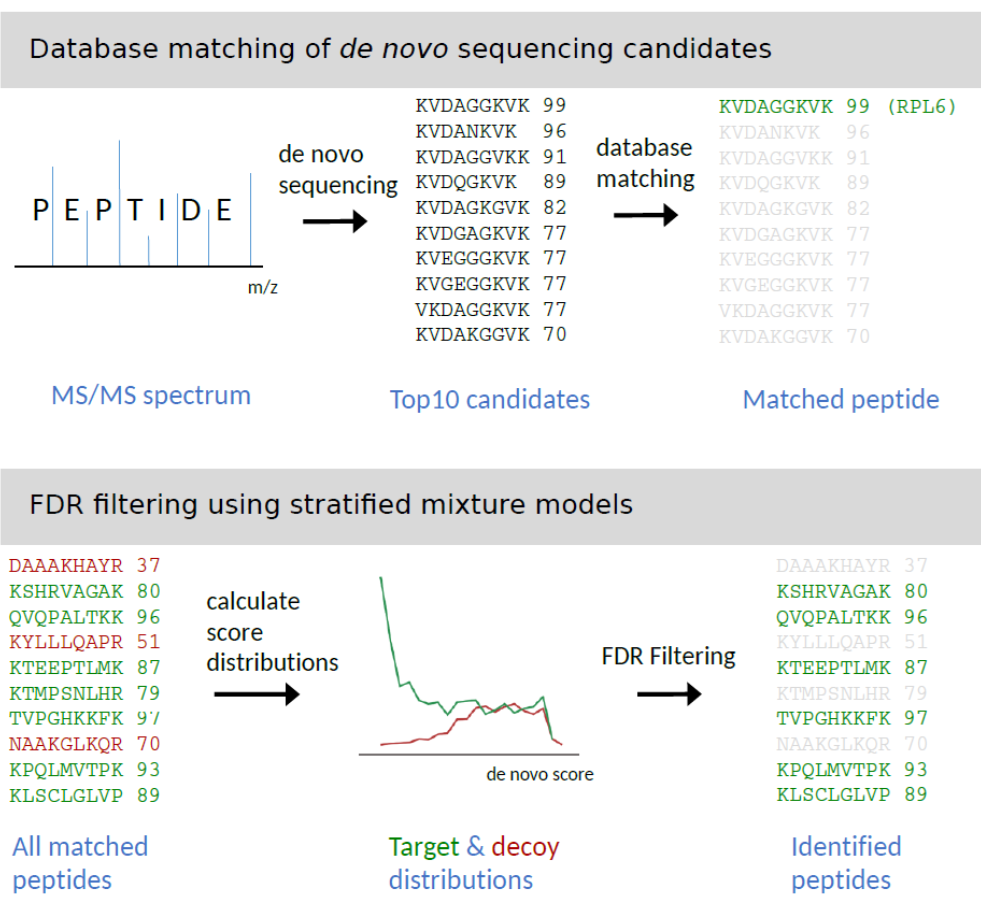
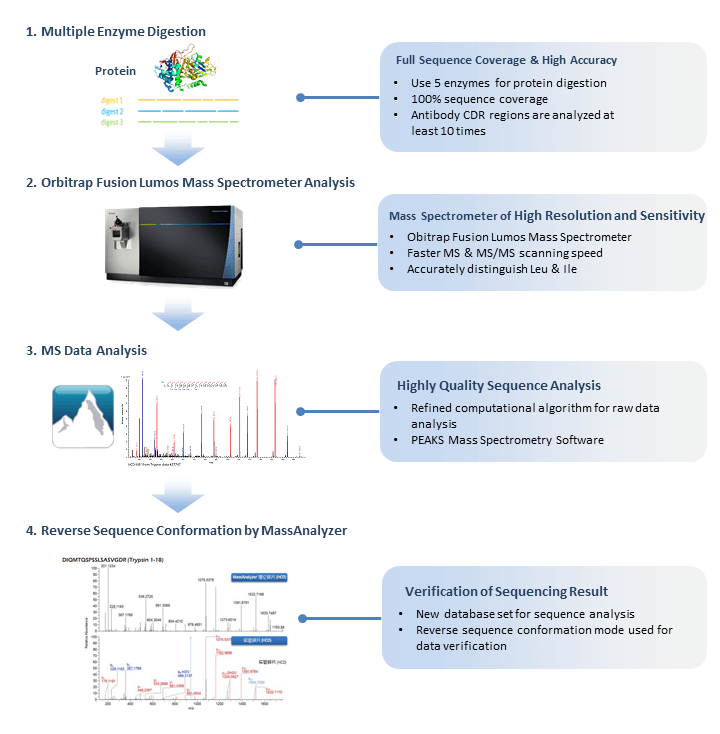
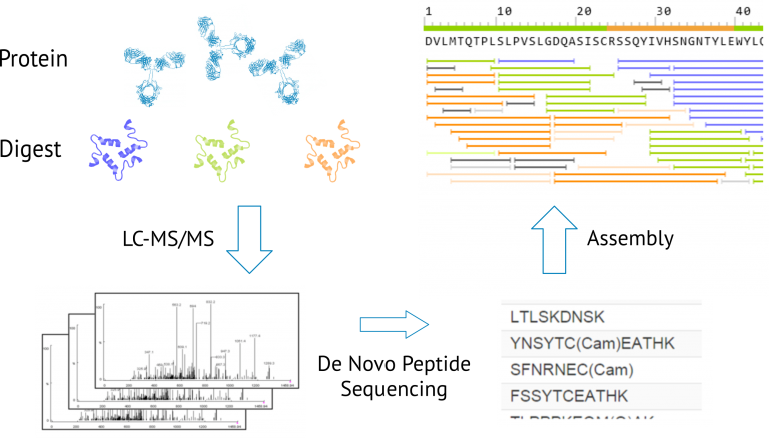
Precision De Novo Peptide Sequencing Using Mirror Proteases of Ac-LysargiNase and Trypsin for Large-scale Proteomics* | Semantic Scholar

Identification and Sequencing of N-Terminal Peptides in Proteins by LC-Fluorescence-MS/MS: An Approach to Replacement of the Edman Degradation | Analytical Chemistry

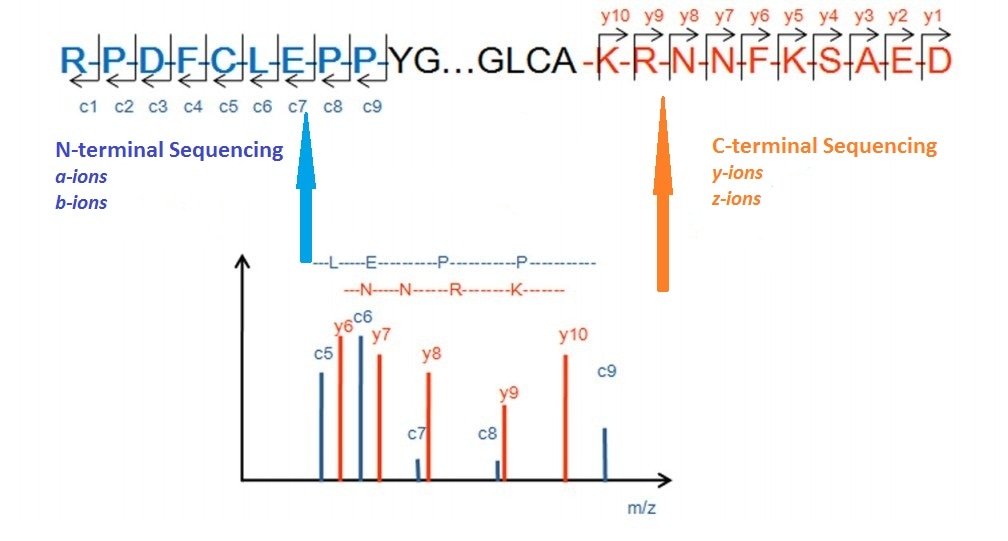
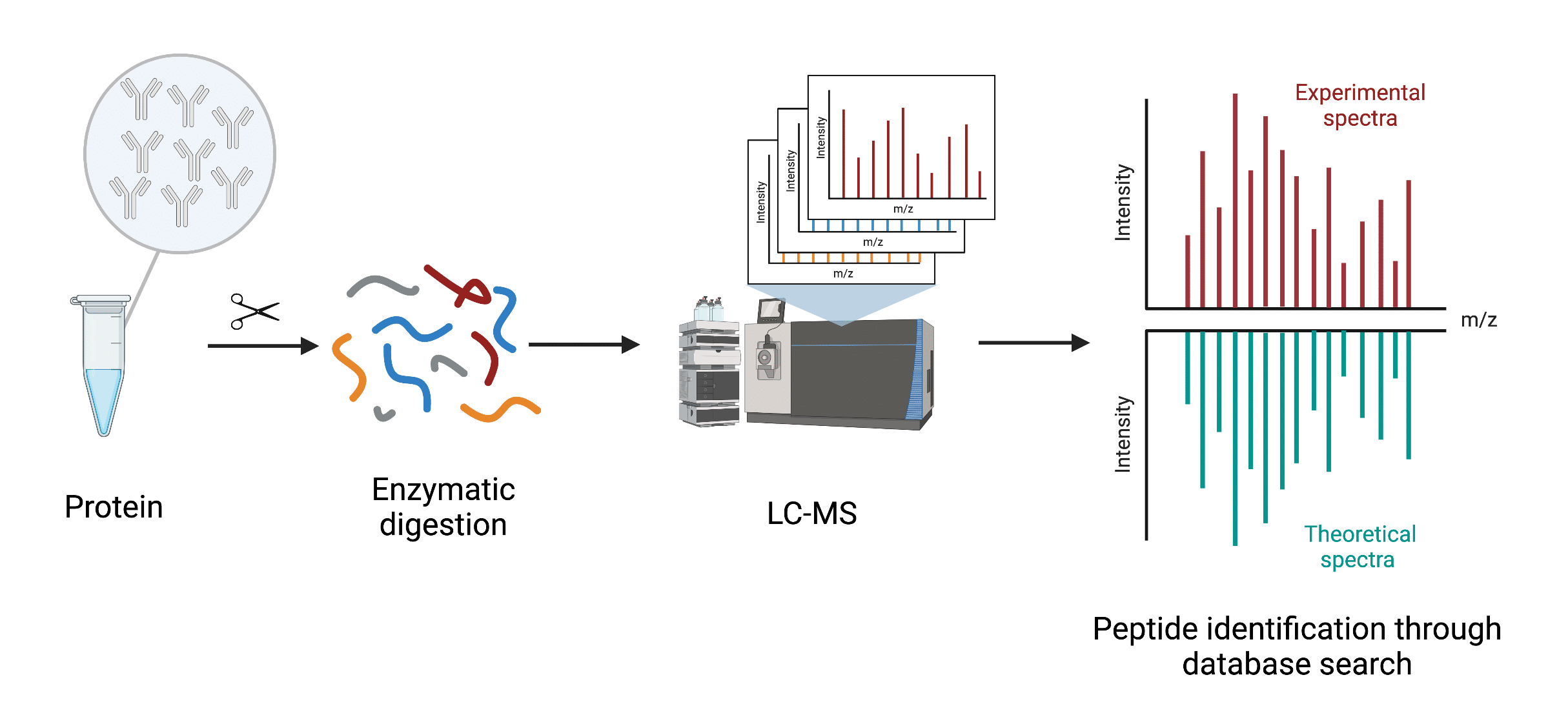
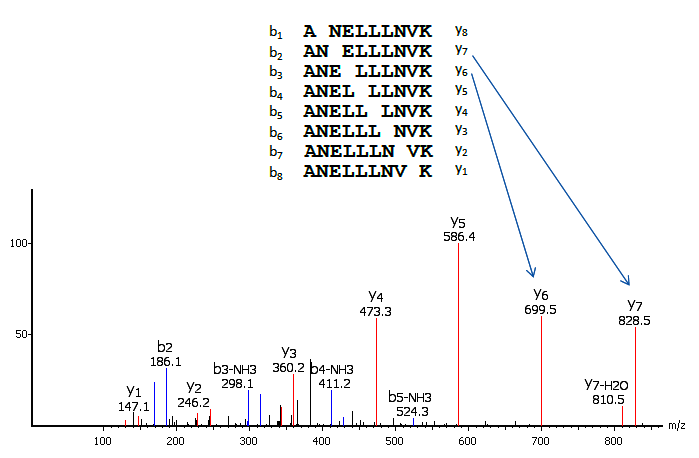
MS/MS for peptide sequencing and identification of oxidative cysteine... | Download Scientific Diagram

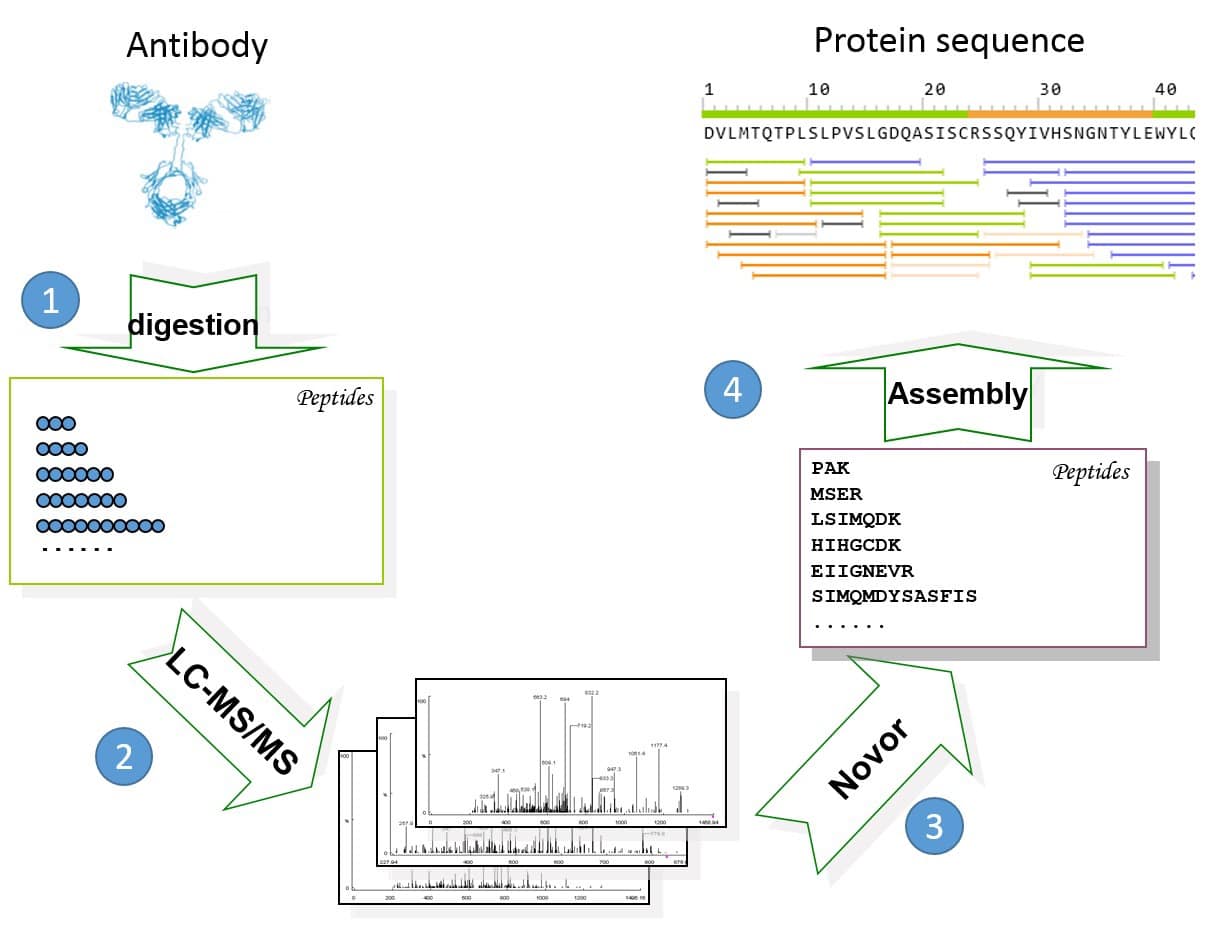
Deep learning enables de novo peptide sequencing from data-independent-acquisition mass spectrometry | Nature Methods
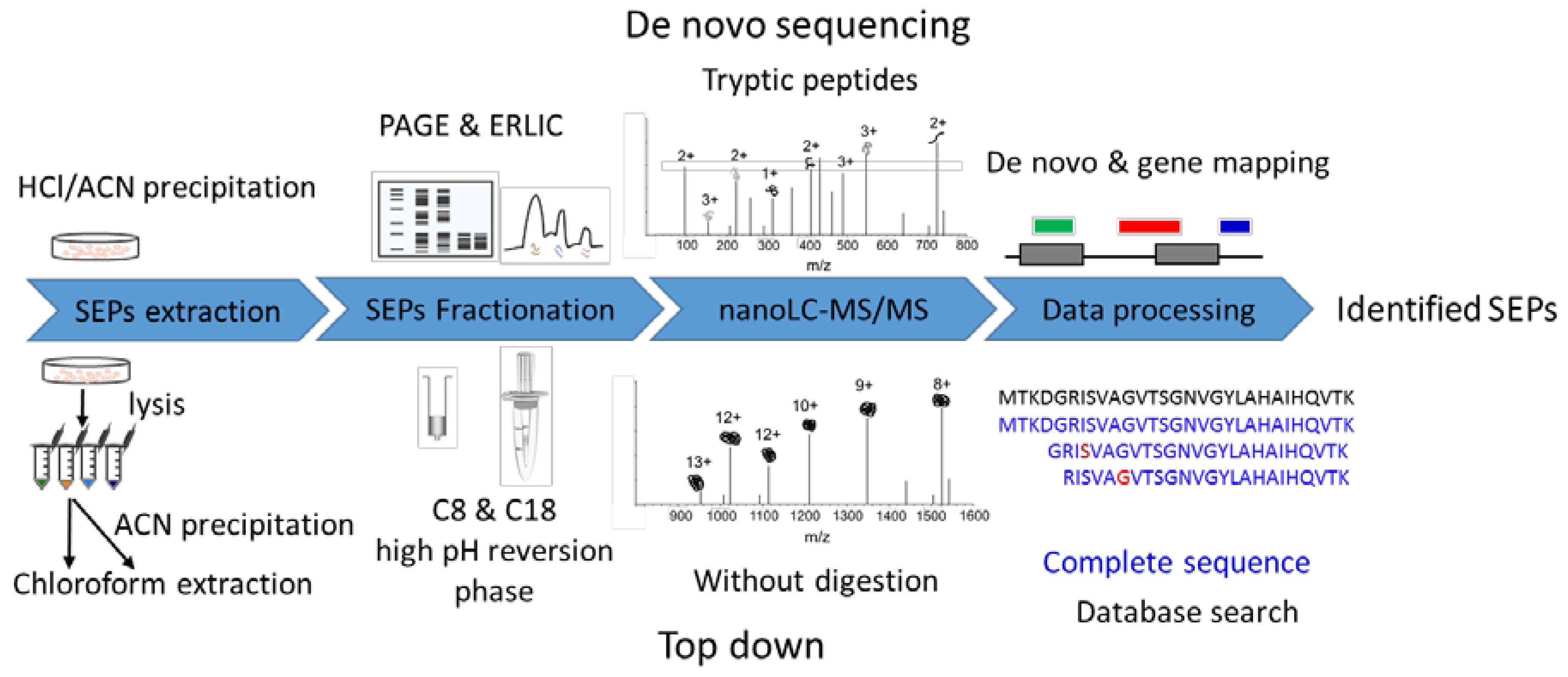
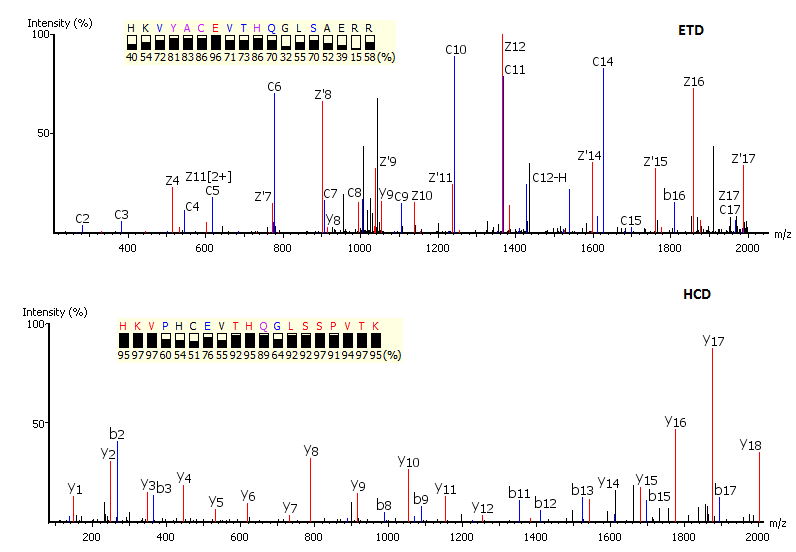
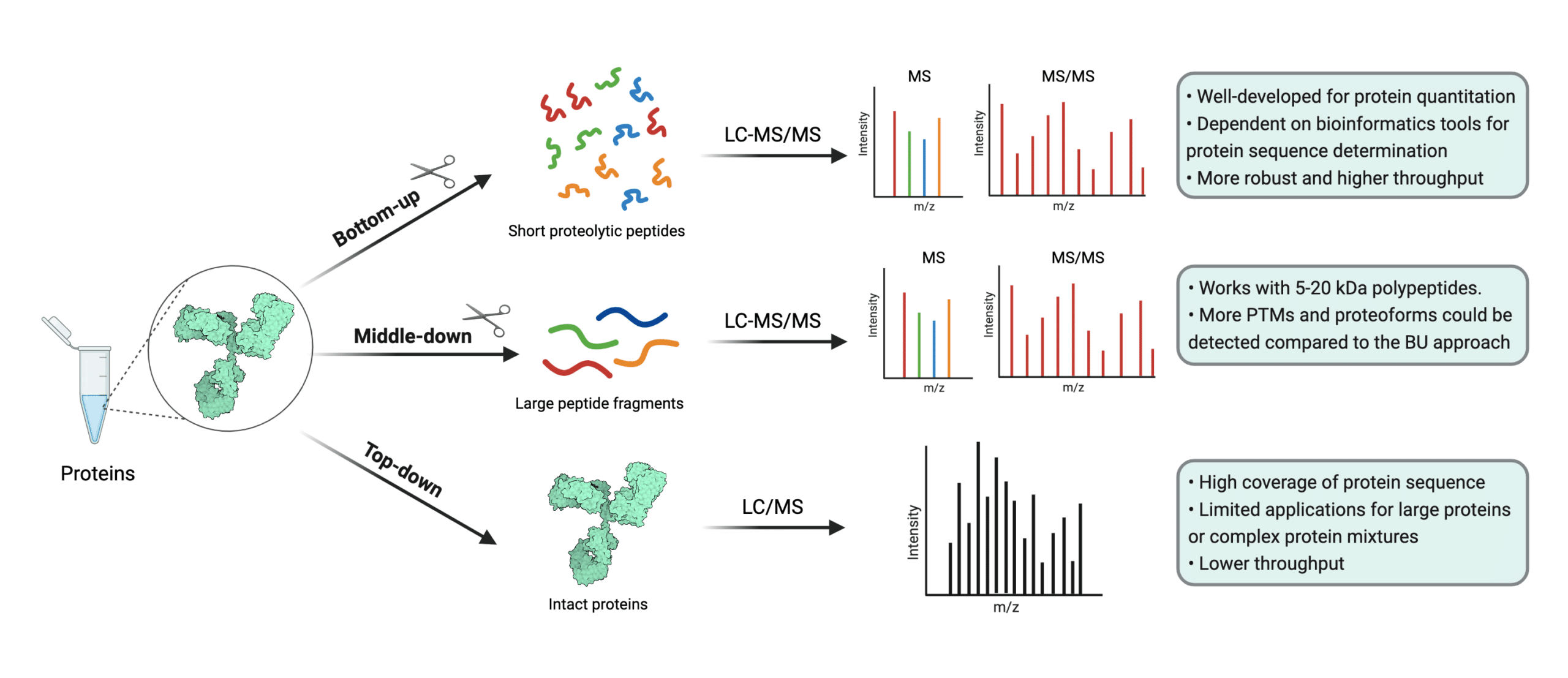
IJMS | Free Full-Text | Improved Identification of Small Open Reading Frames Encoded Peptides by Top-Down Proteomic Approaches and De Novo Sequencing