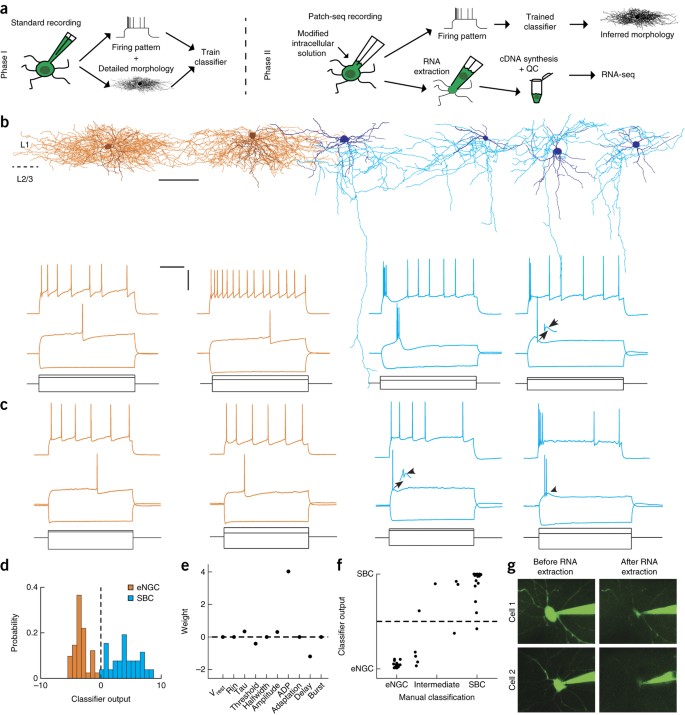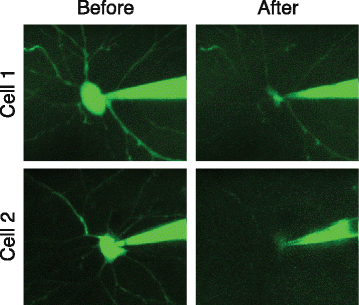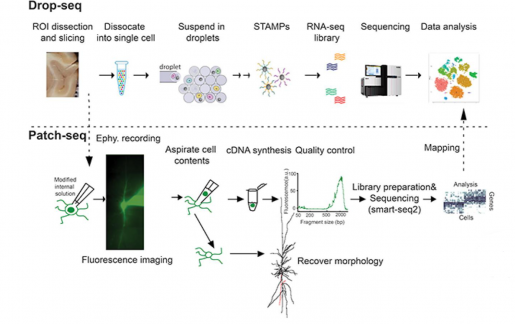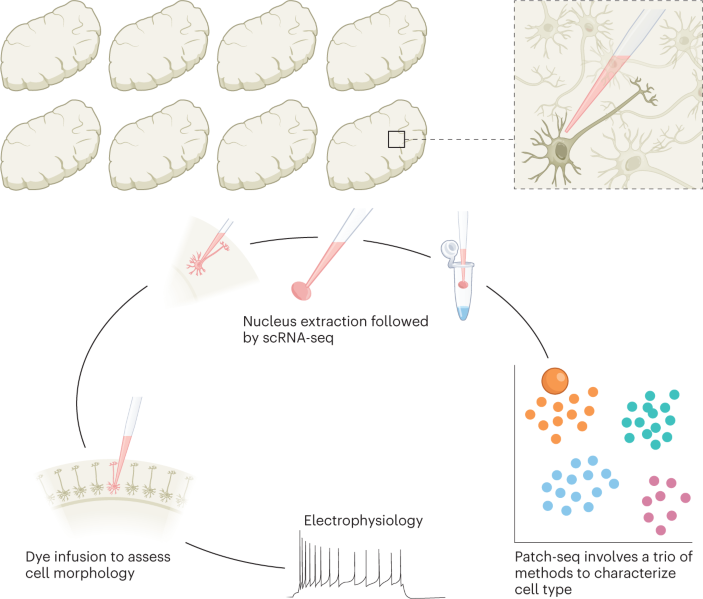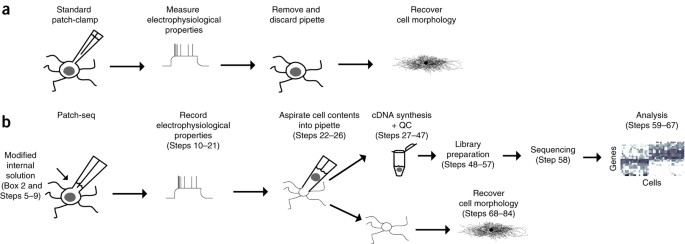
Multimodal profiling of single-cell morphology, electrophysiology, and gene expression using Patch-seq | Nature Protocols
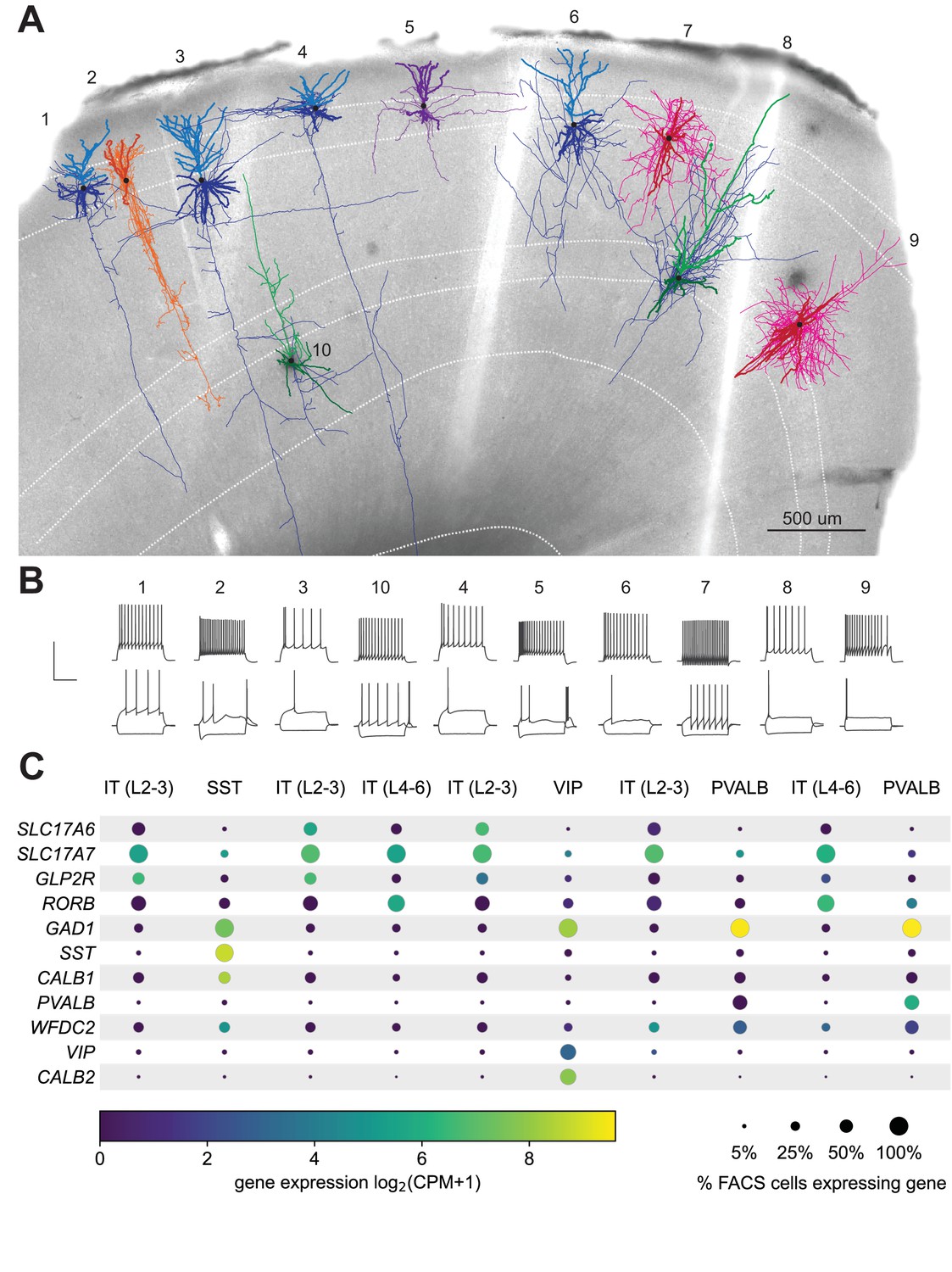
Scaled, high fidelity electrophysiological, morphological, and transcriptomic cell characterization | eLife
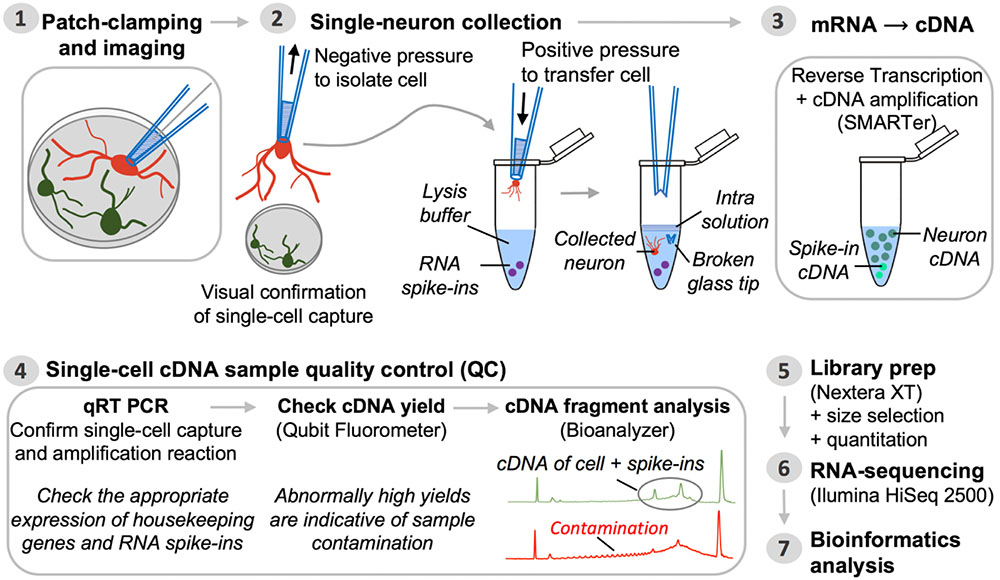
Frontiers | Patch-Seq Protocol to Analyze the Electrophysiology, Morphology and Transcriptome of Whole Single Neurons Derived From Human Pluripotent Stem Cells
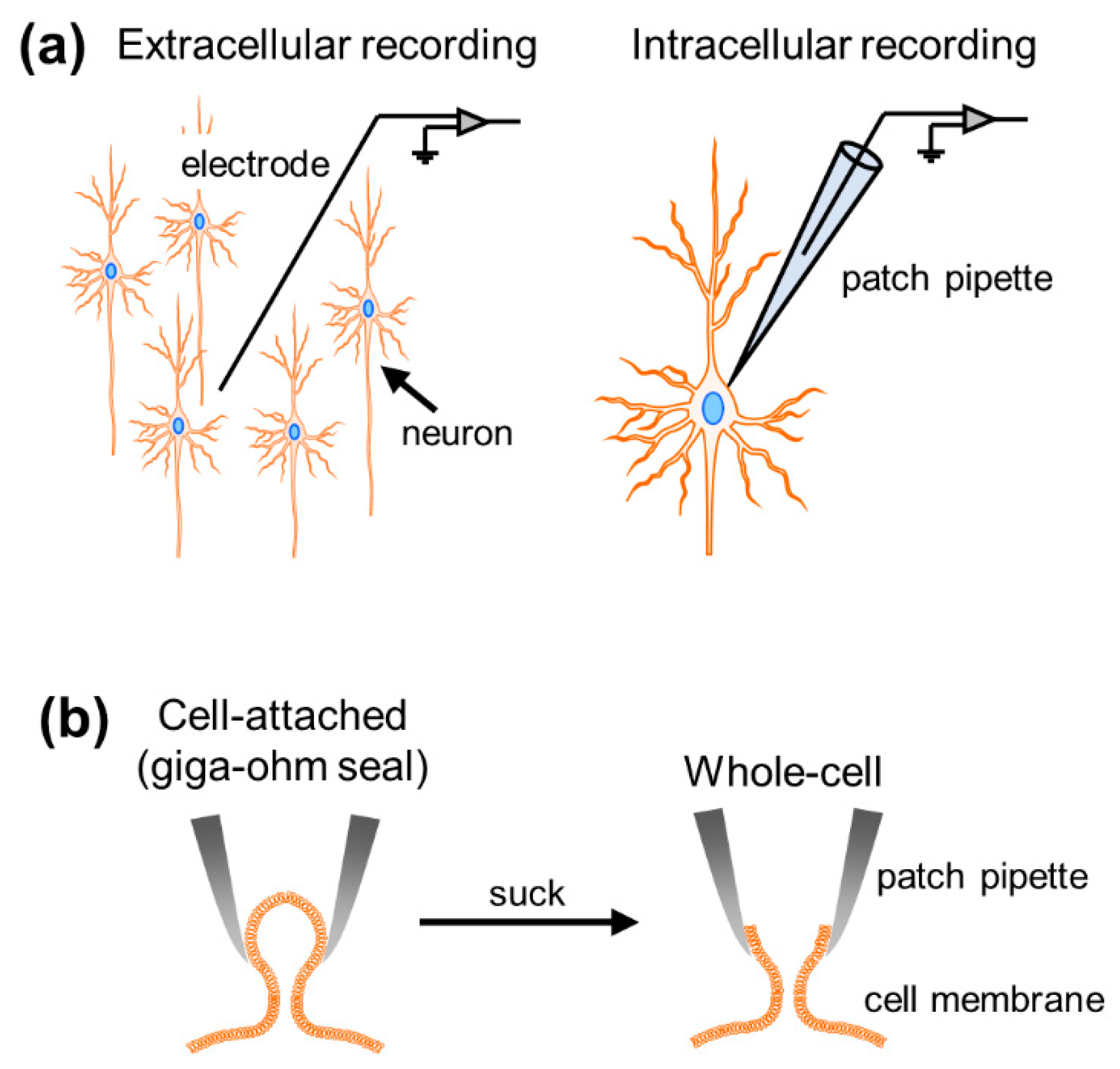
Sensors | Free Full-Text | In Vivo Whole-Cell Patch-Clamp Methods: Recent Technical Progress and Future Perspectives

Felix Ritort Lab | Seminar “RNA sequencing technologies for health and disease monitoring: single-cell patch-seq in diabetes and cell-free RNA in fetal-maternal health”

Patch-sequencing of the subiculum-projecting VIP+ cells. a Schematic of... | Download Scientific Diagram
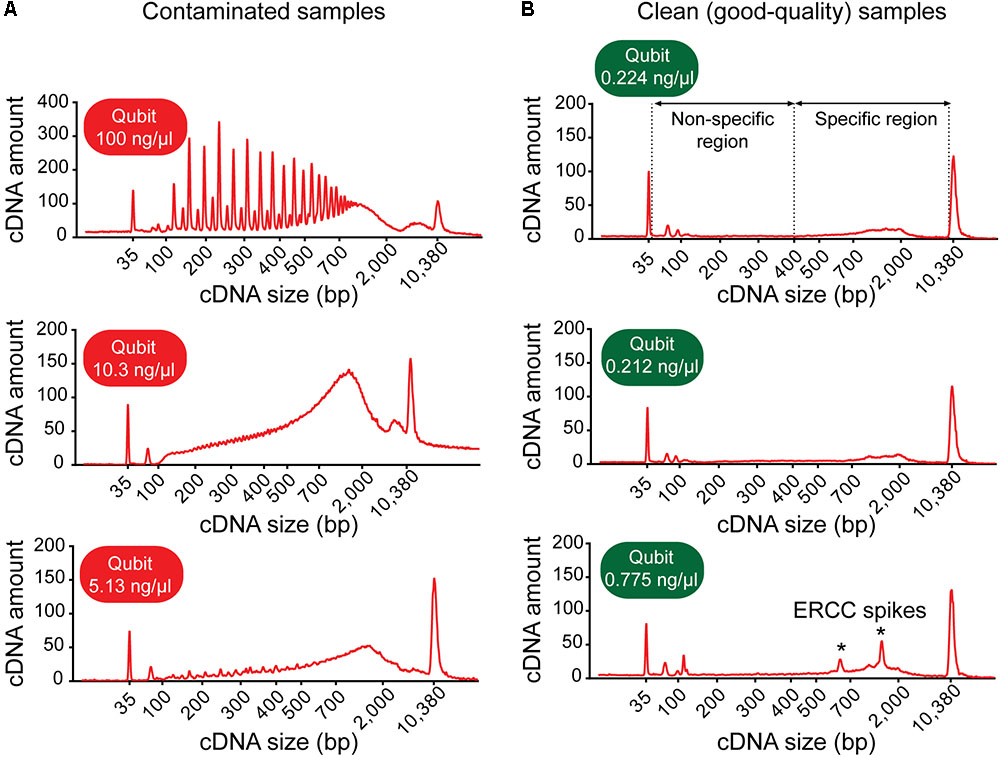
Frontiers | Patch-Seq Protocol to Analyze the Electrophysiology, Morphology and Transcriptome of Whole Single Neurons Derived From Human Pluripotent Stem Cells

High fidelity electrophysiological, morphological, and transcriptomic cell characterization using a refined Patch-seq protocol | bioRxiv
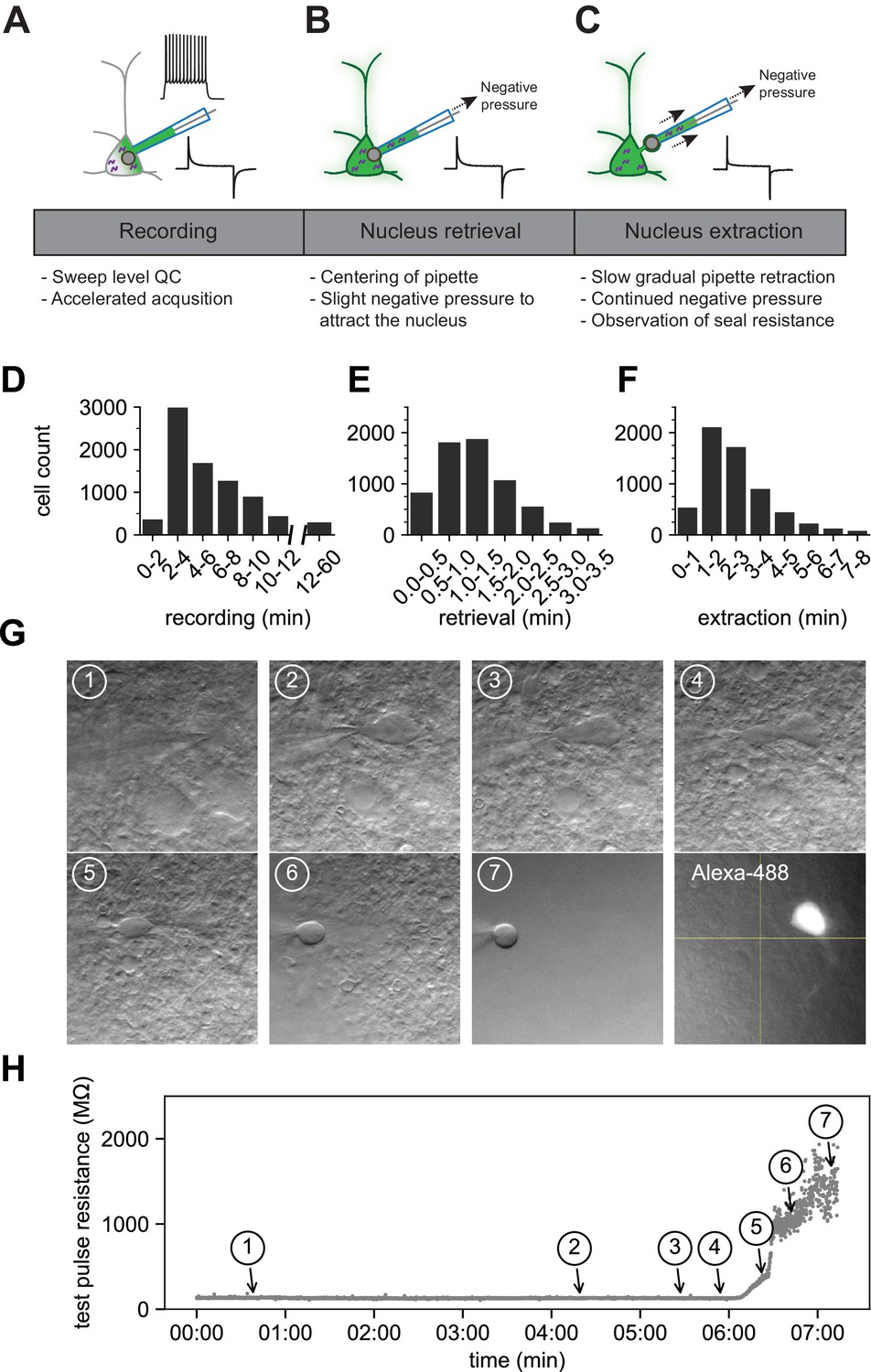
Scaled, high fidelity electrophysiological, morphological, and transcriptomic cell characterization | eLife

RADAR-seq: A RAre DAmage and Repair sequencing method for detecting DNA damage on a genome-wide scale - ScienceDirect