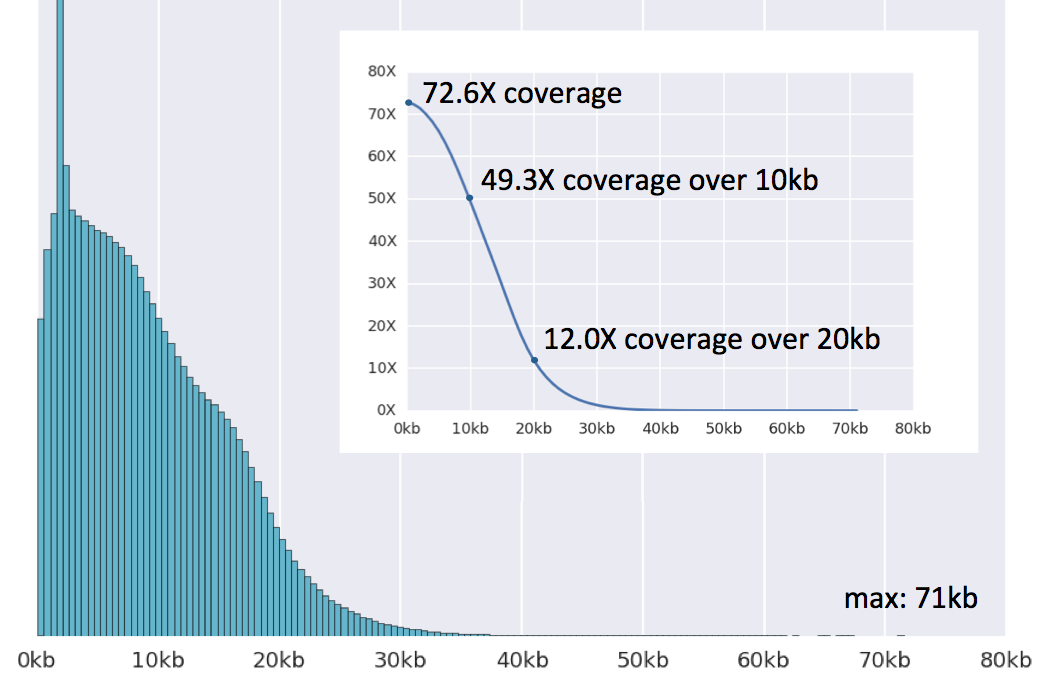
PacBio single molecule long-read sequencing provides insight into the complexity and diversity of the Pinctada fucata martensii transcriptome | BMC Genomics | Full Text

PacBio single-molecule long-read sequencing of C. porrectum. (a) Reads... | Download Scientific Diagram
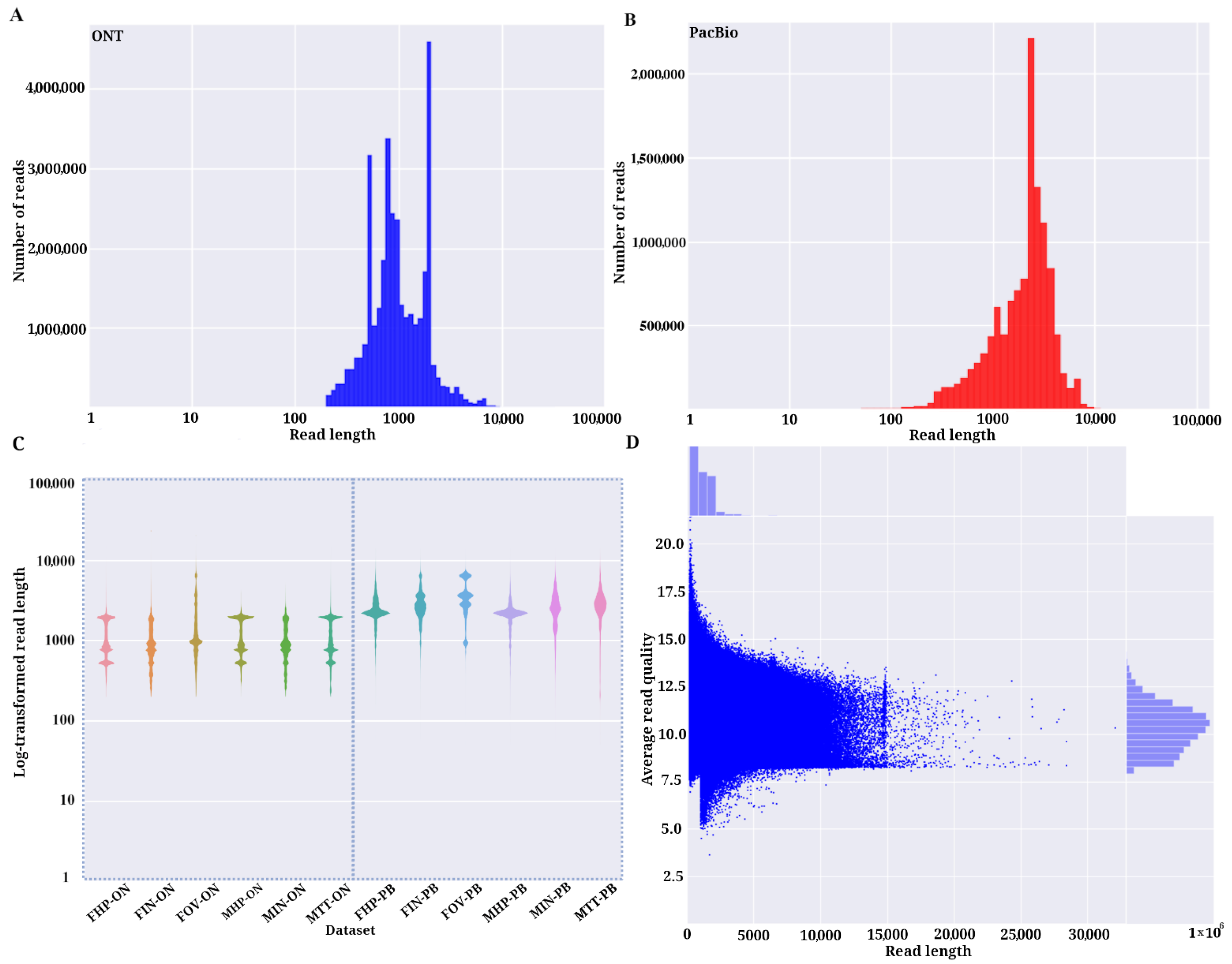
Life | Free Full-Text | Comparative Analysis of PacBio and Oxford Nanopore Sequencing Technologies for Transcriptomic Landscape Identification of Penaeus monodon
![SMRT sequencing of the full-length transcriptome of the Rhynchophorus ferrugineus (Coleoptera: Curculionidae) [PeerJ] SMRT sequencing of the full-length transcriptome of the Rhynchophorus ferrugineus (Coleoptera: Curculionidae) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/9133/1/fig-1-full.png)
SMRT sequencing of the full-length transcriptome of the Rhynchophorus ferrugineus (Coleoptera: Curculionidae) [PeerJ]

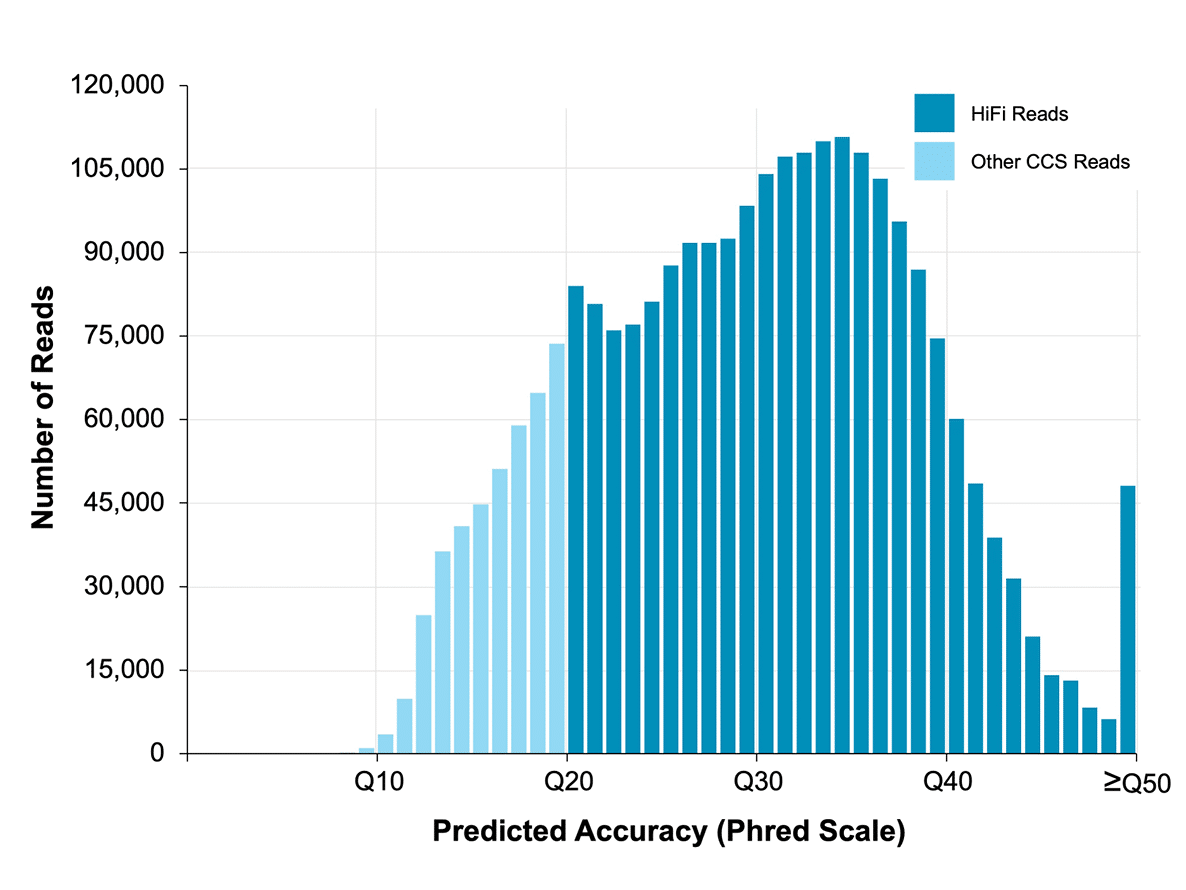
Improved metagenome assemblies and taxonomic binning using long-read circular consensus sequence data | Scientific Reports

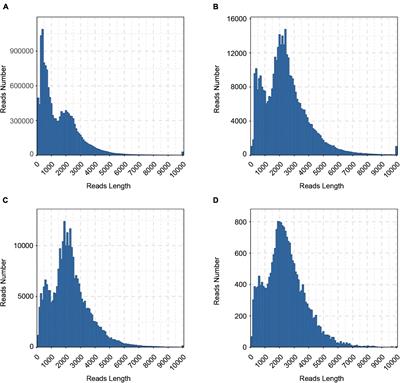
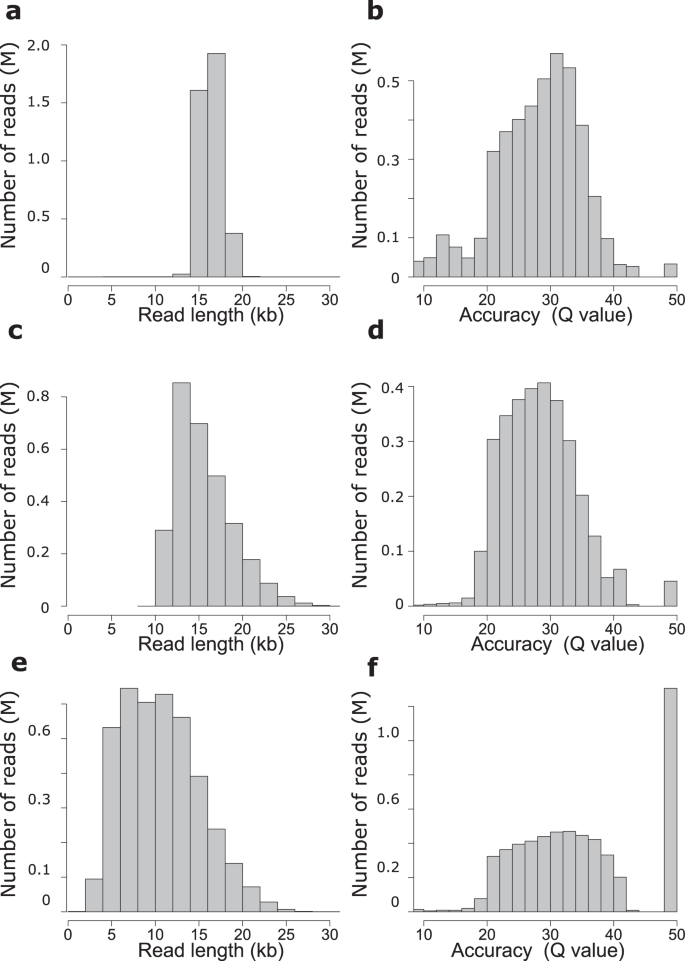
PacBio read length and quality distribution. The top and bottom left... | Download Scientific Diagram
![SMRT sequencing of the full-length transcriptome of the Rhynchophorus ferrugineus (Coleoptera: Curculionidae) [PeerJ] SMRT sequencing of the full-length transcriptome of the Rhynchophorus ferrugineus (Coleoptera: Curculionidae) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/9133/1/fig-11-full.png)
SMRT sequencing of the full-length transcriptome of the Rhynchophorus ferrugineus (Coleoptera: Curculionidae) [PeerJ]
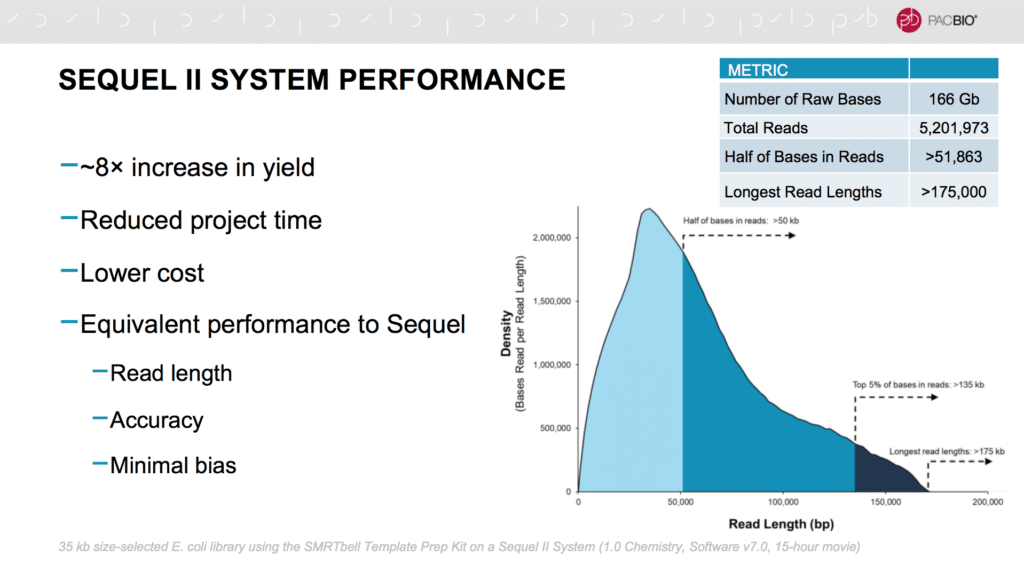
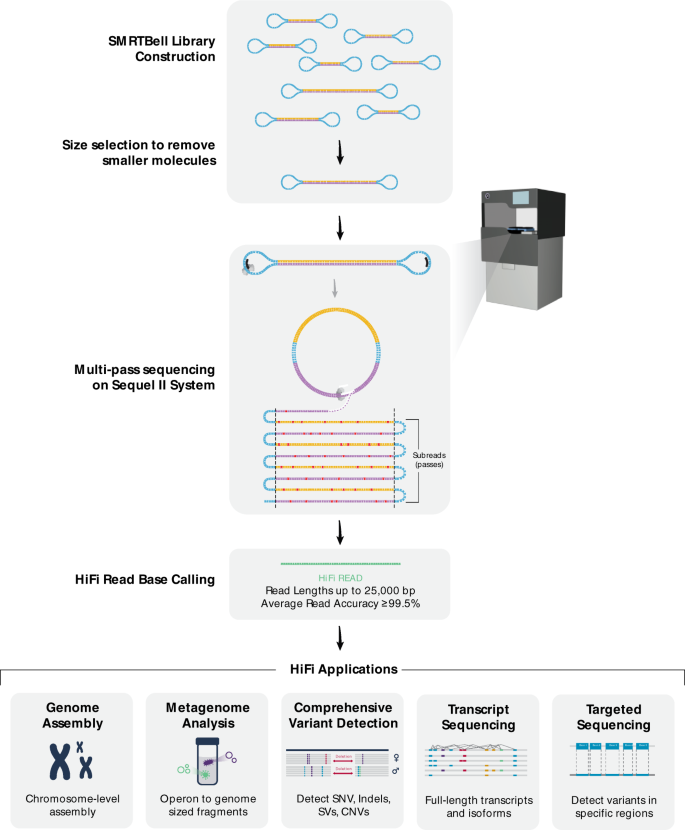
Users Report on SMRT Sequencing: Sequel II System, HiFi Reads, Iso-Seq Analysis, Microbiomes, and More - PacBio

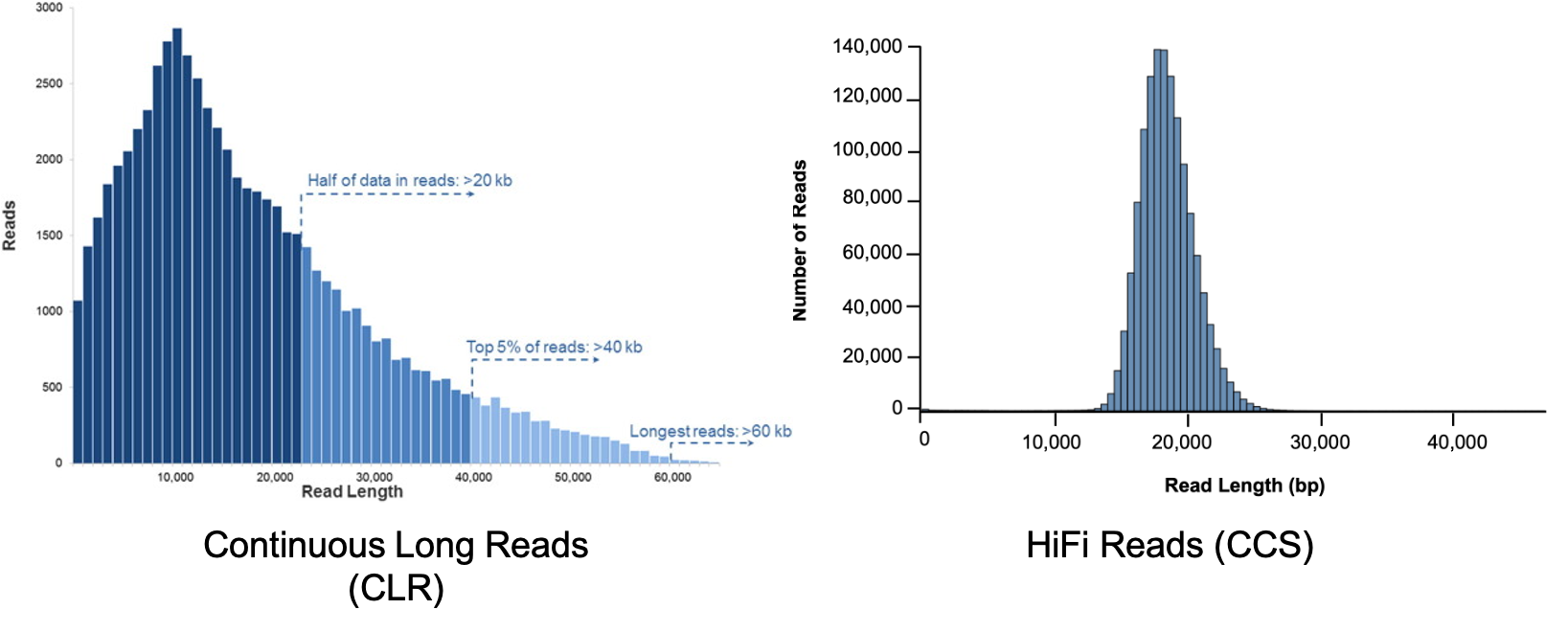
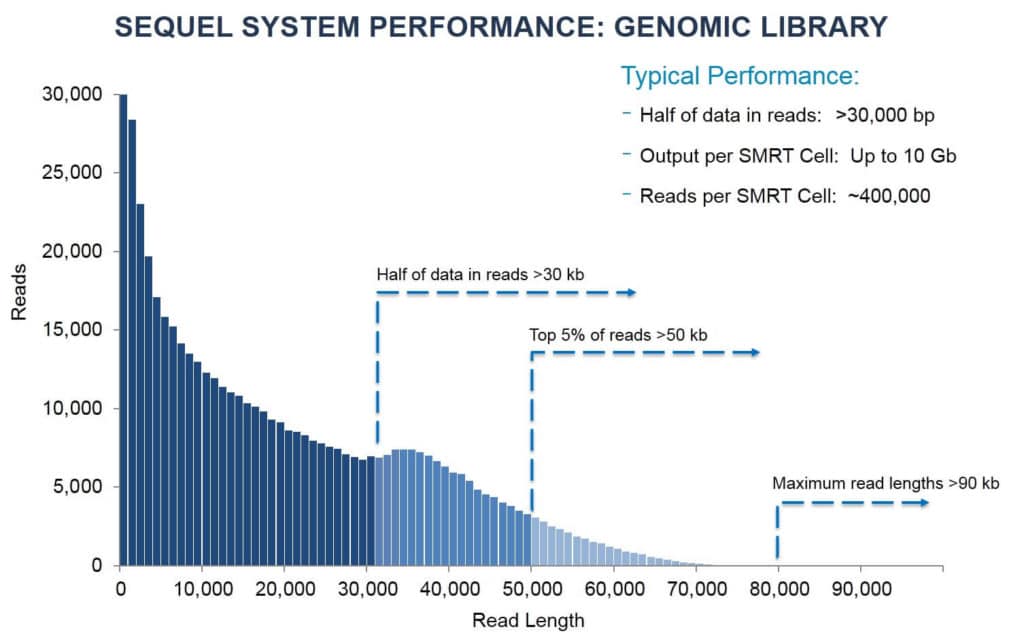
PacBio long-and HiFi-sequenced read length distribution vs. read count... | Download Scientific Diagram

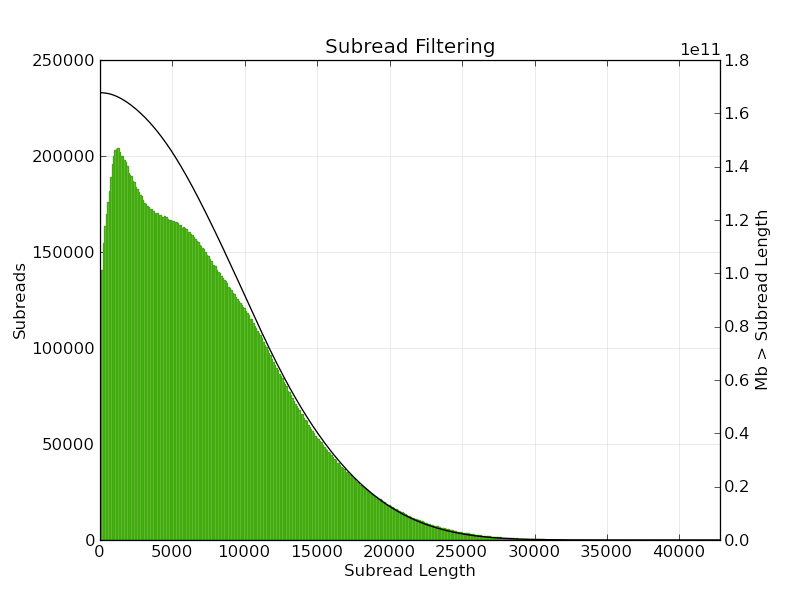
PacBio sequencing parameters. a Read length distribution of the 17,802... | Download Scientific Diagram