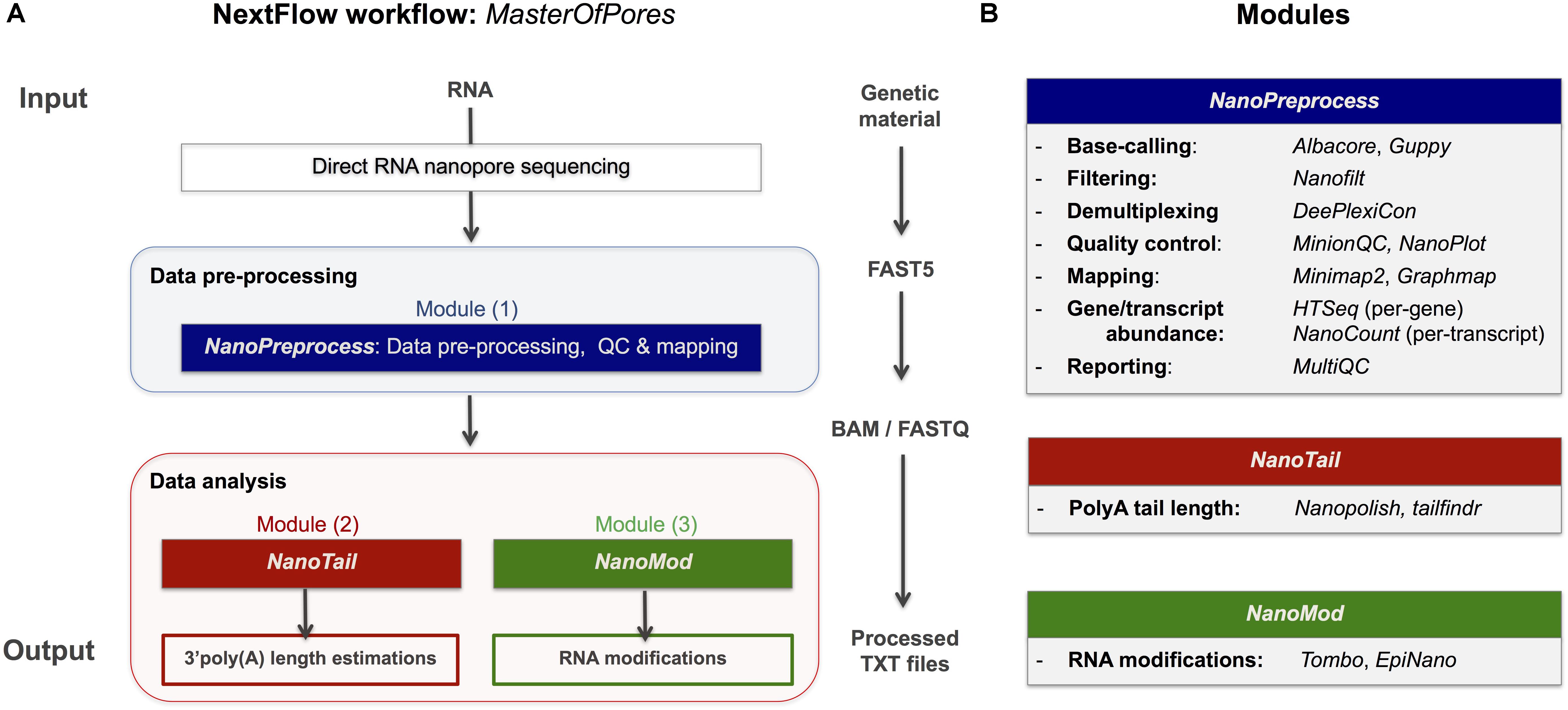
Frontiers | MasterOfPores: A Workflow for the Analysis of Oxford Nanopore Direct RNA Sequencing Datasets

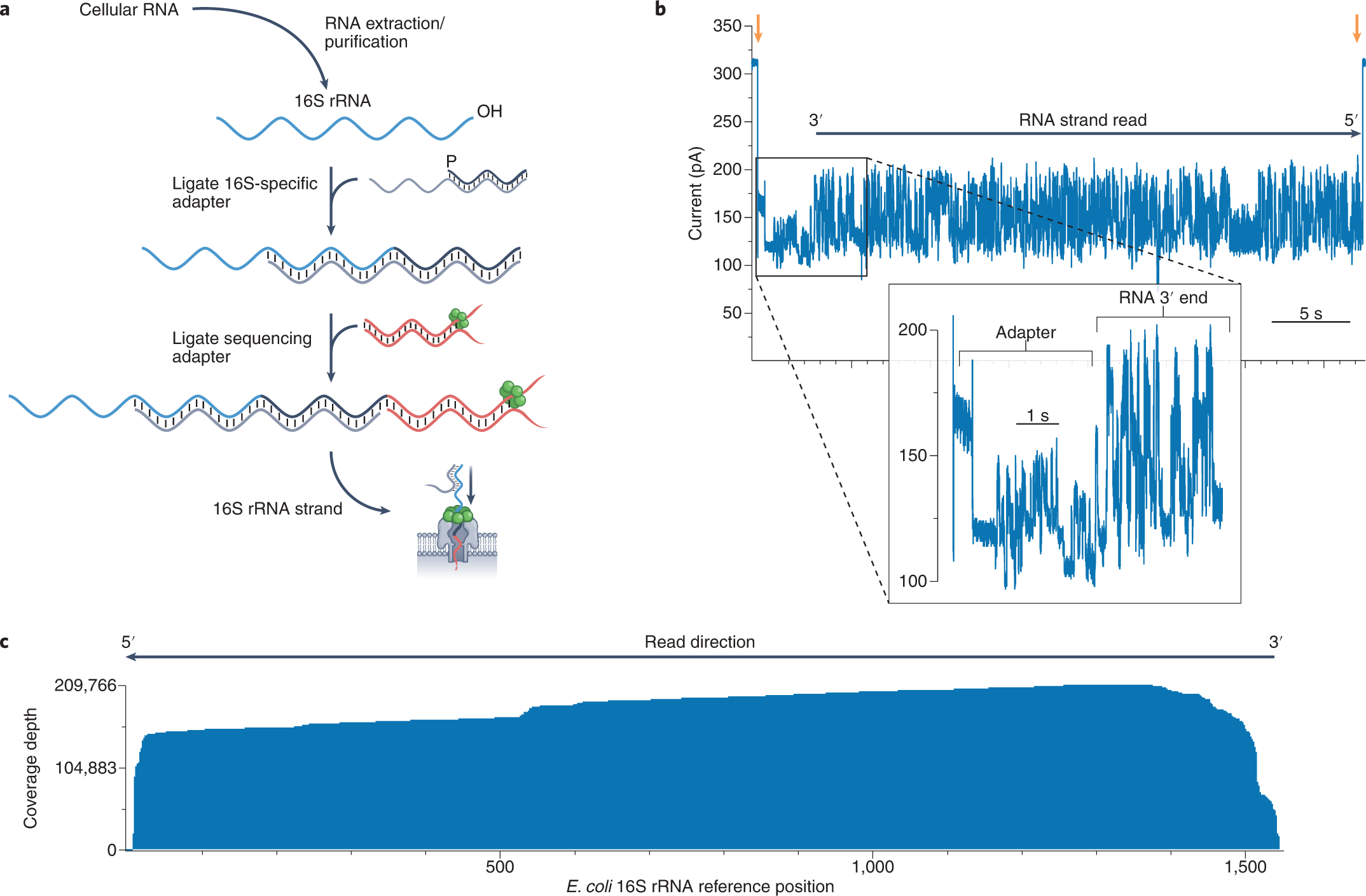
Nanopore-based native RNA sequencing provides insights into prokaryotic transcription, operon structures, rRNA maturation and modifications | bioRxiv

Beyond sequencing: machine learning algorithms extract biology hidden in Nanopore signal data: Trends in Genetics

Applications and potentials of nanopore sequencing in the (epi)genome and (epi)transcriptome era - ScienceDirect

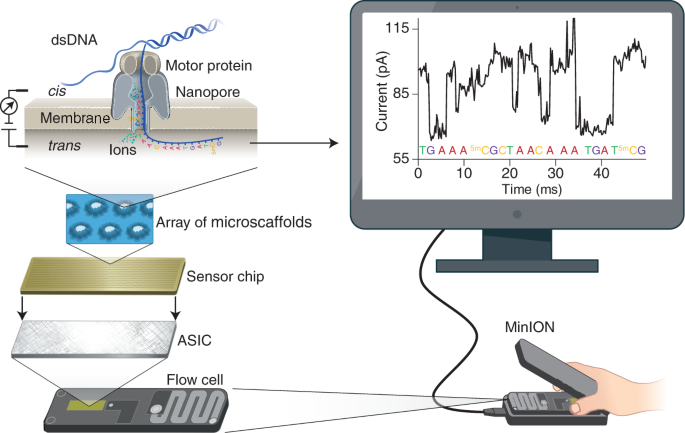
Oxford Nanopore on Twitter: "On the cover of @naturemethods, Read "Highly parallel direct RNA sequencing on an array of nanopores" https://t.co/Pk9a6czgy0 https://t.co/DBuyuupqld" / Twitter
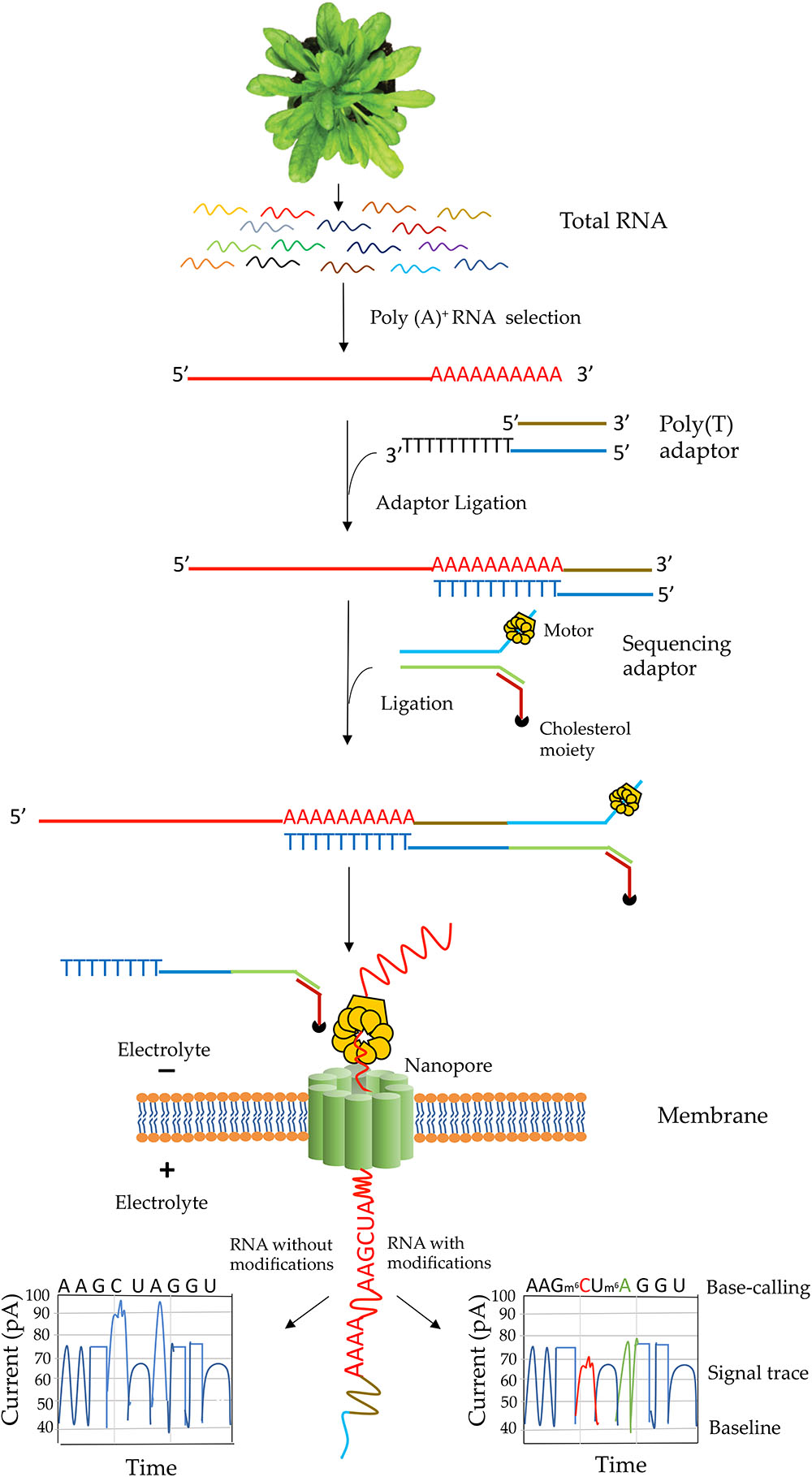
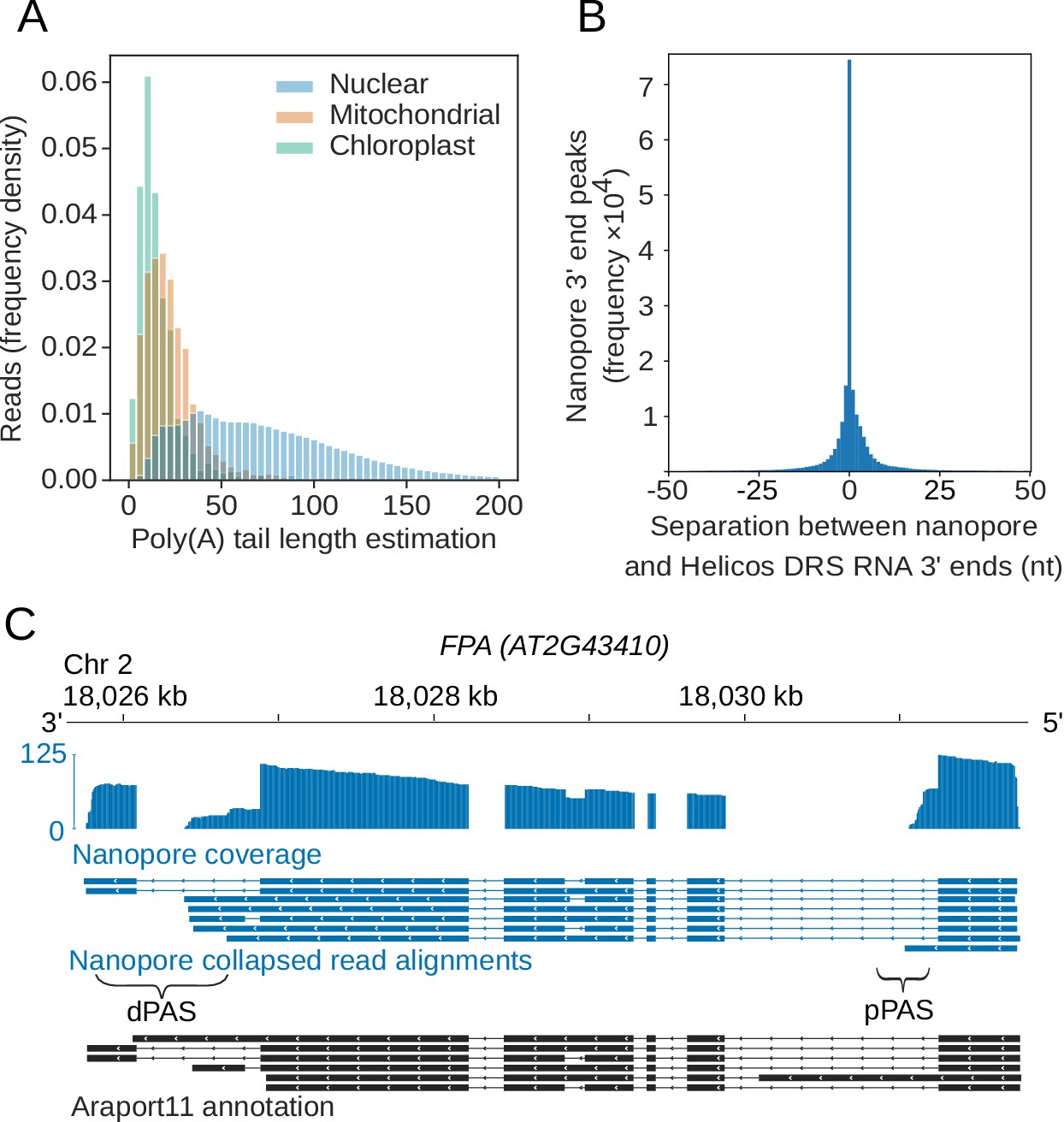
Frontiers | Analysis of Transcriptome and Epitranscriptome in Plants Using PacBio Iso-Seq and Nanopore-Based Direct RNA Sequencing

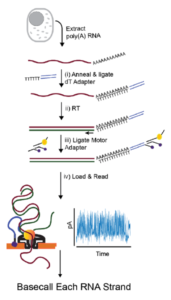
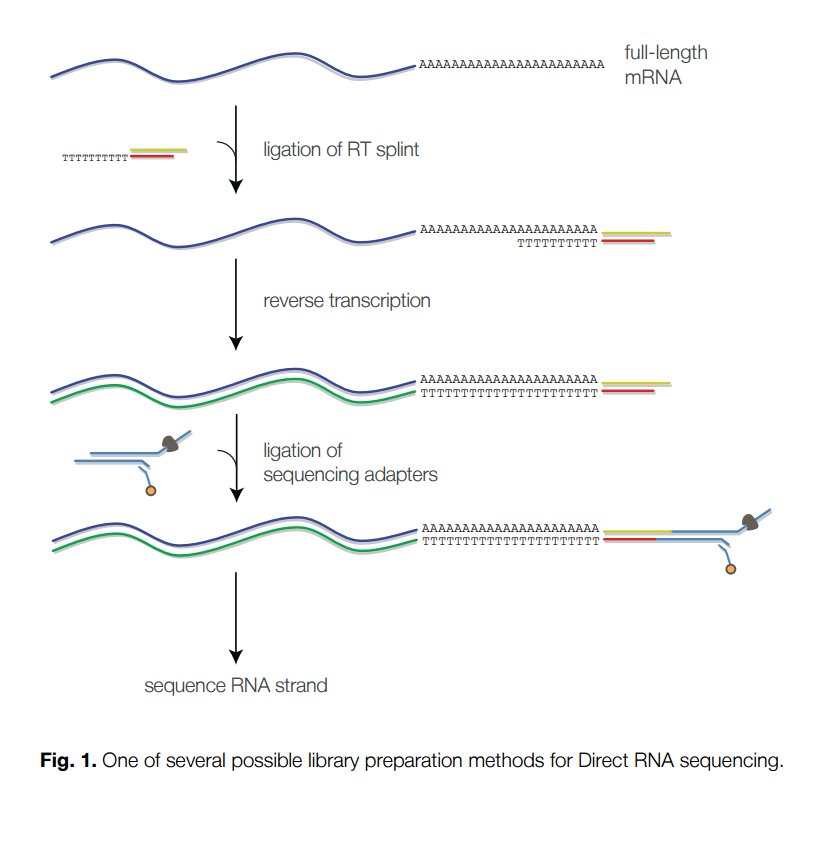
Direct RNA sequencing library preparation steps using Oxford Nanopore... | Download Scientific Diagram

Direct detection of RNA modifications and structure using single molecule nanopore sequencing | bioRxiv

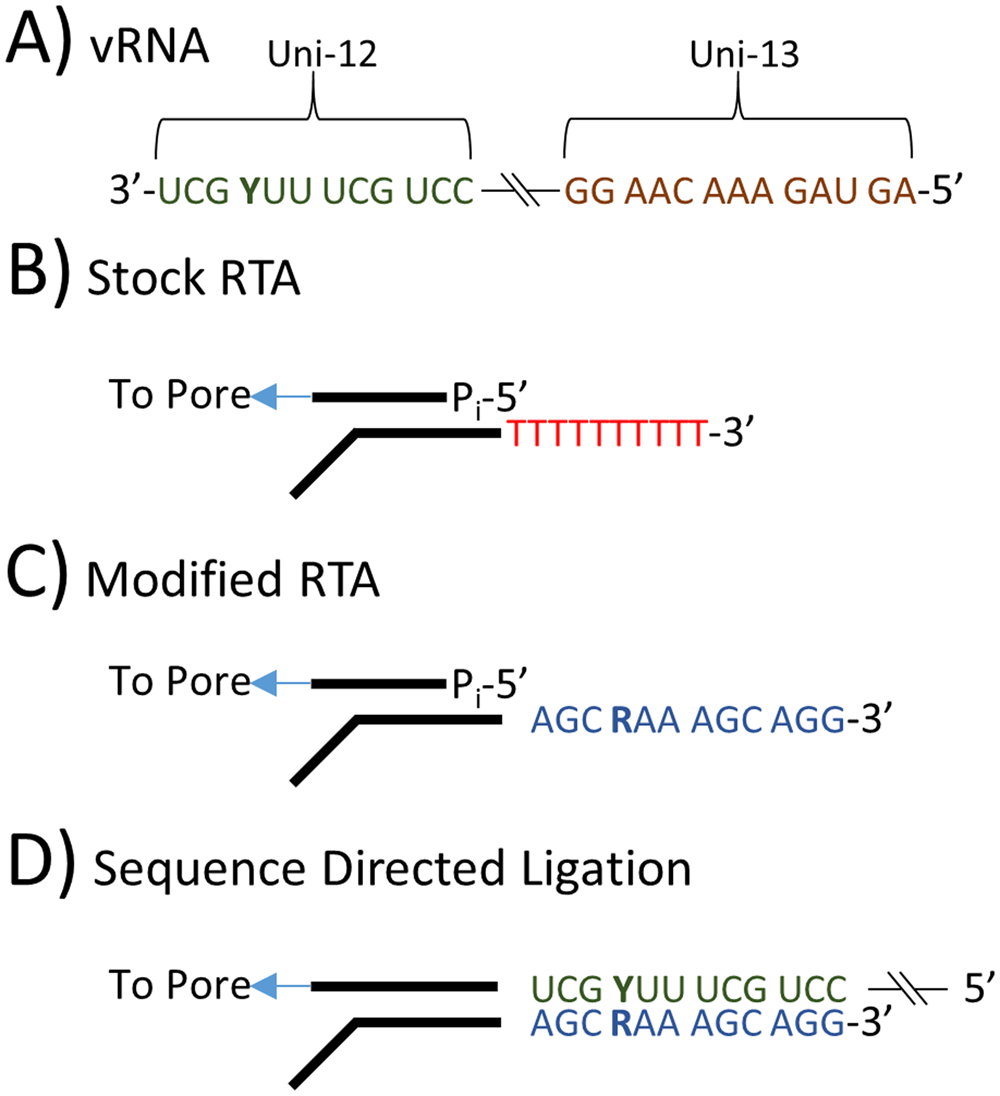
Toxins | Free Full-Text | RIPpore: A Novel Host-Derived Method for the Identification of Ricin Intoxication through Oxford Nanopore Direct RNA Sequencing
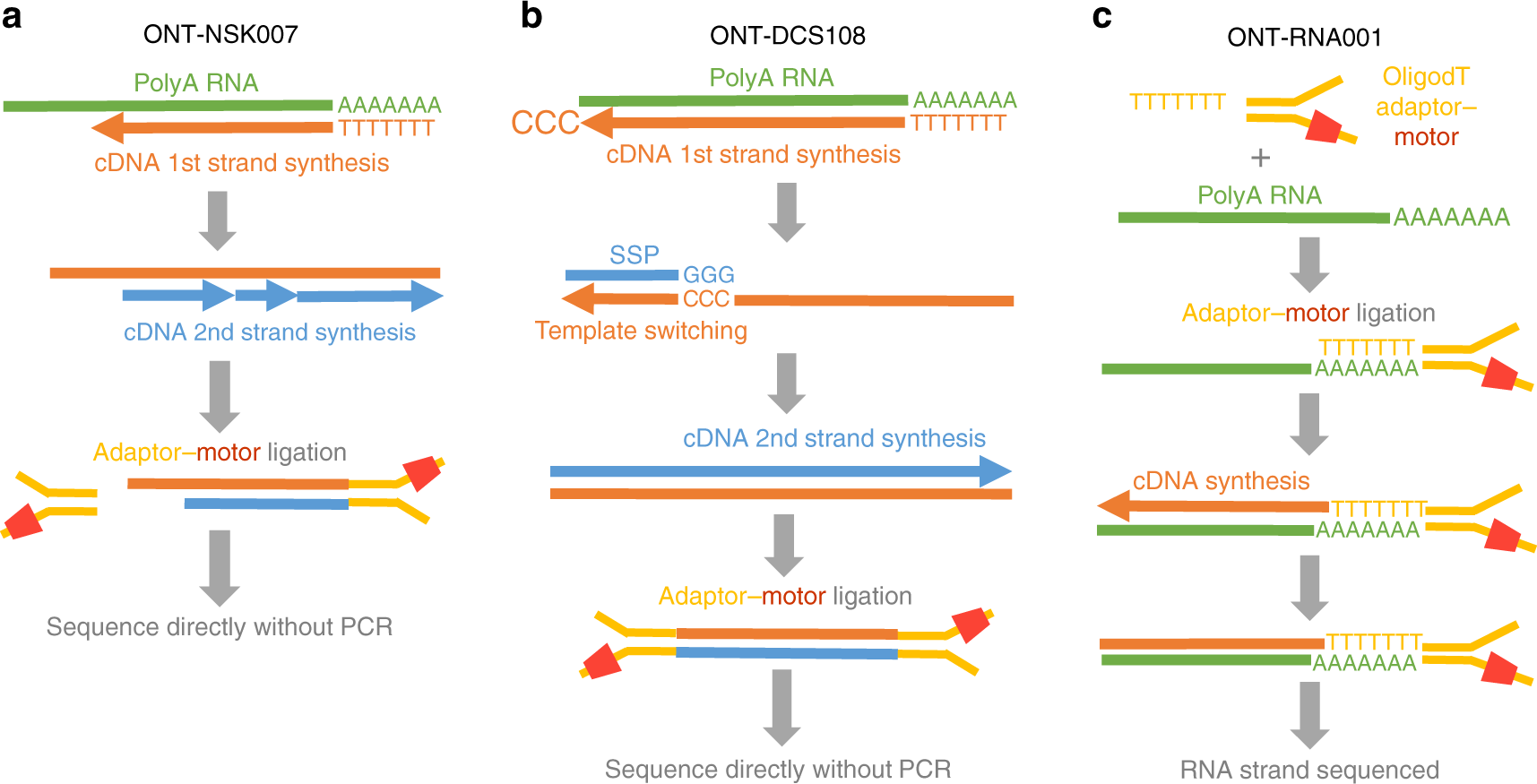
A comprehensive examination of Nanopore native RNA sequencing for characterization of complex transcriptomes | Nature Communications
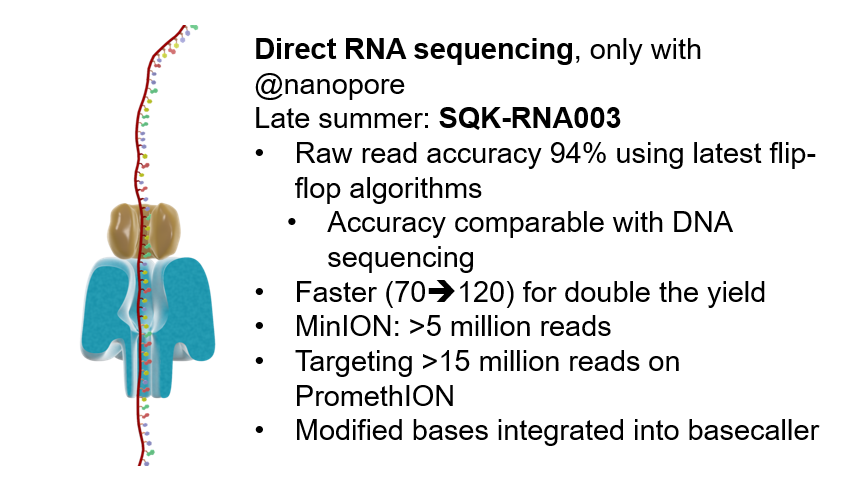

Oxford Nanopore on Twitter: "AH: Direct RNA with @nanopore will get closer to mainstream method this year #nanoporeconf https://t.co/3LrUy8fGog" / Twitter