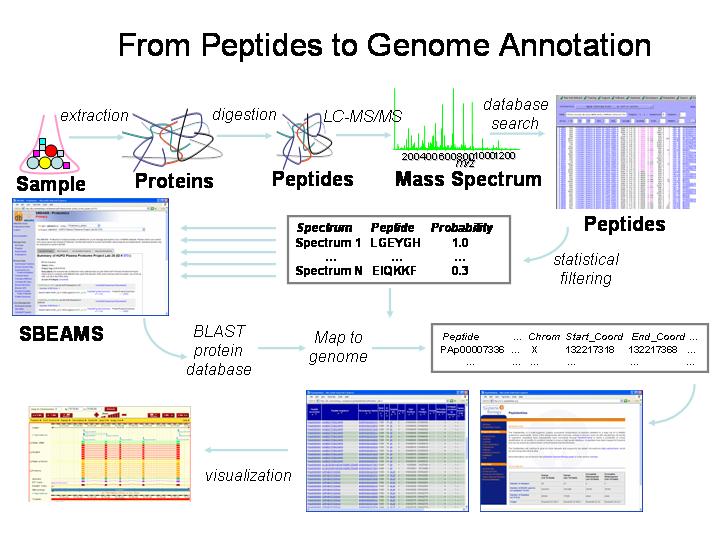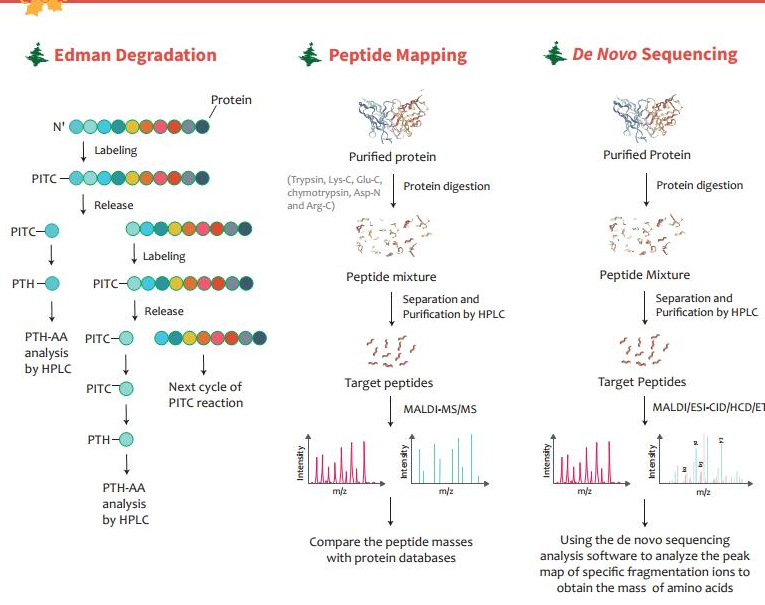
Creative Proteomics on Twitter: "Chose suitable protein sequencing methods for your project This infographic introduces three protein sequencing methods: Edman degradation, peptide mapping, and de novo protein sequencing. https://t.co/kmI8xFuhlW https ...

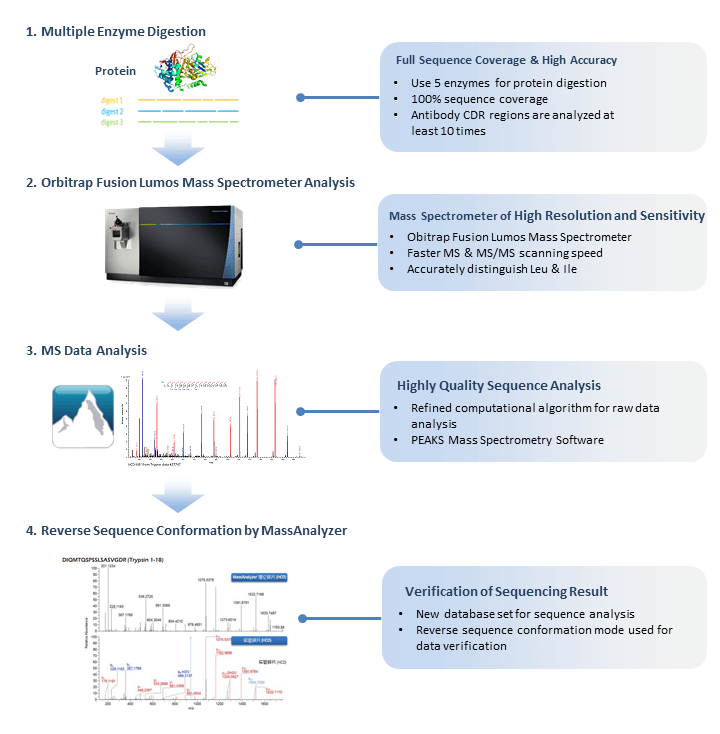
Identification and Sequencing of N-Terminal Peptides in Proteins by LC-Fluorescence-MS/MS: An Approach to Replacement of the Edman Degradation | Analytical Chemistry
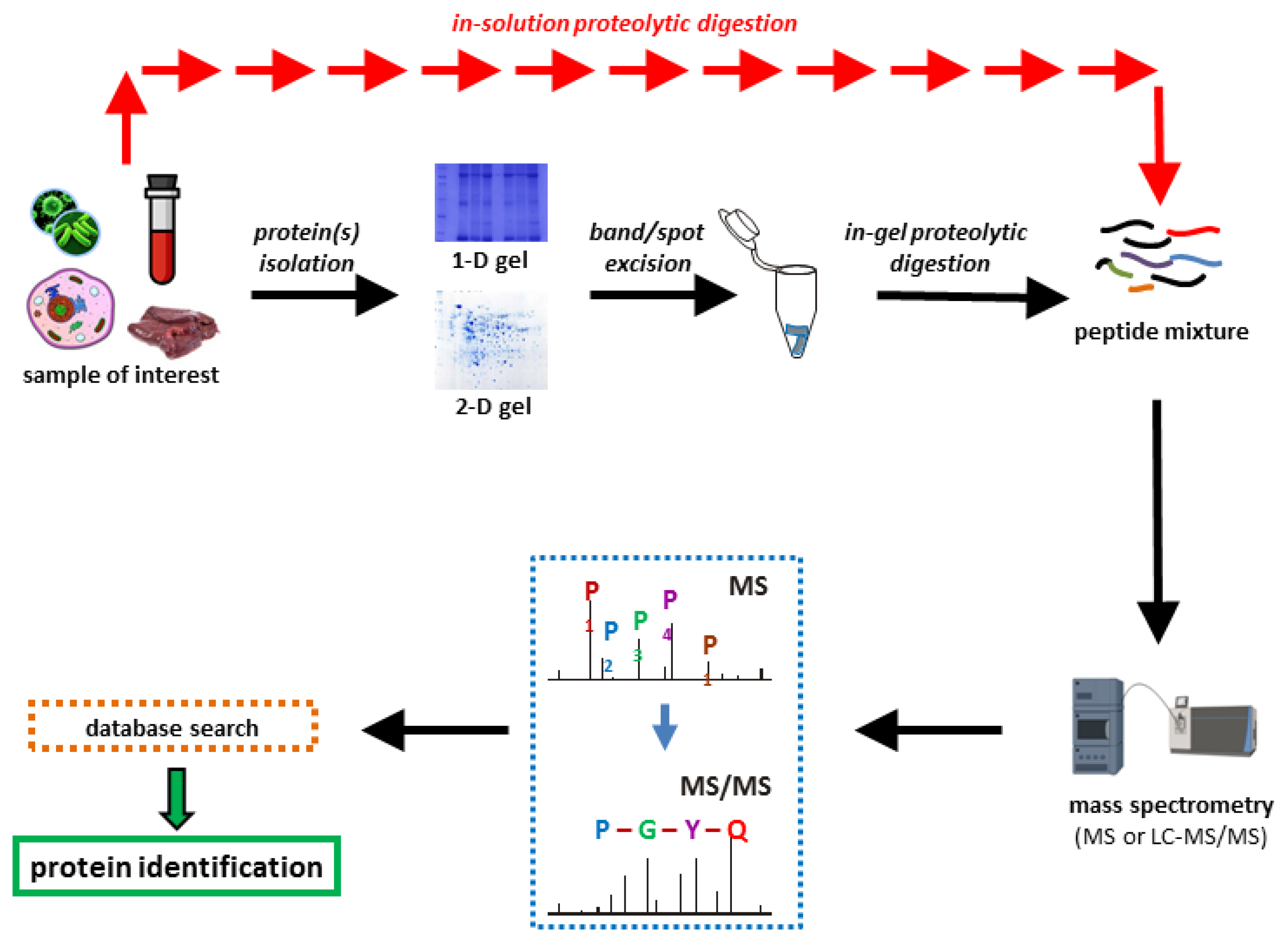
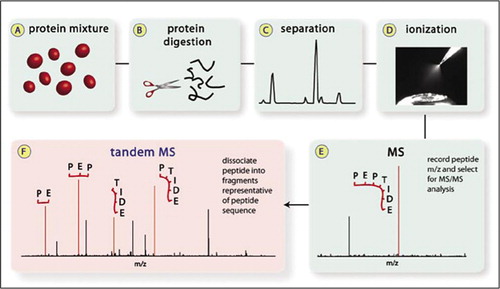
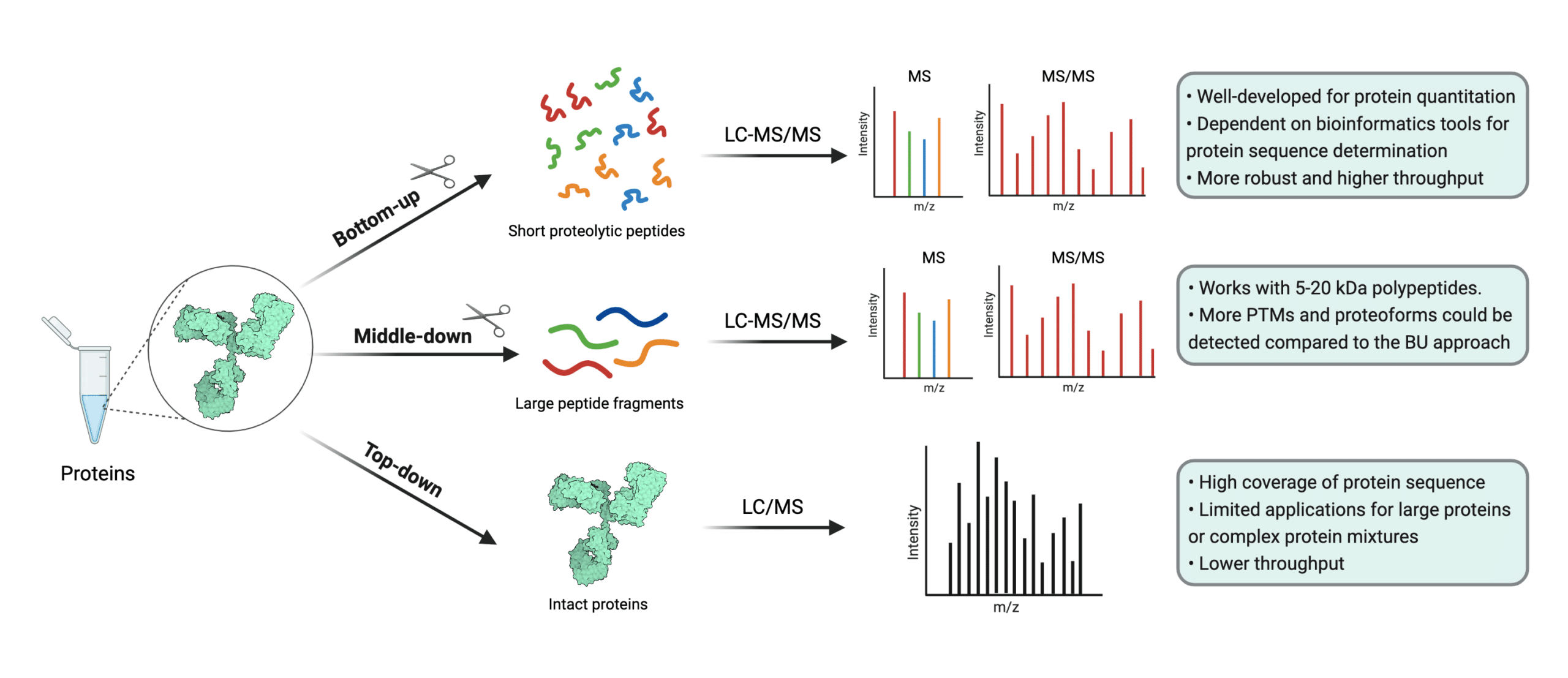
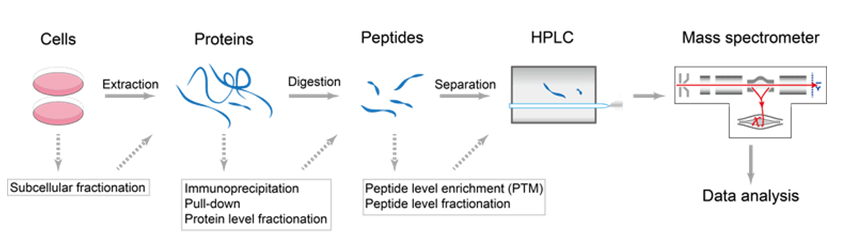
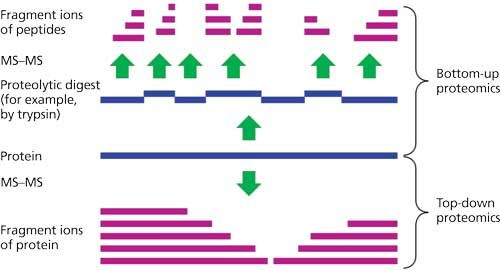
Proteomes | Free Full-Text | A Critical Review of Bottom-Up Proteomics: The Good, the Bad, and the Future of This Field

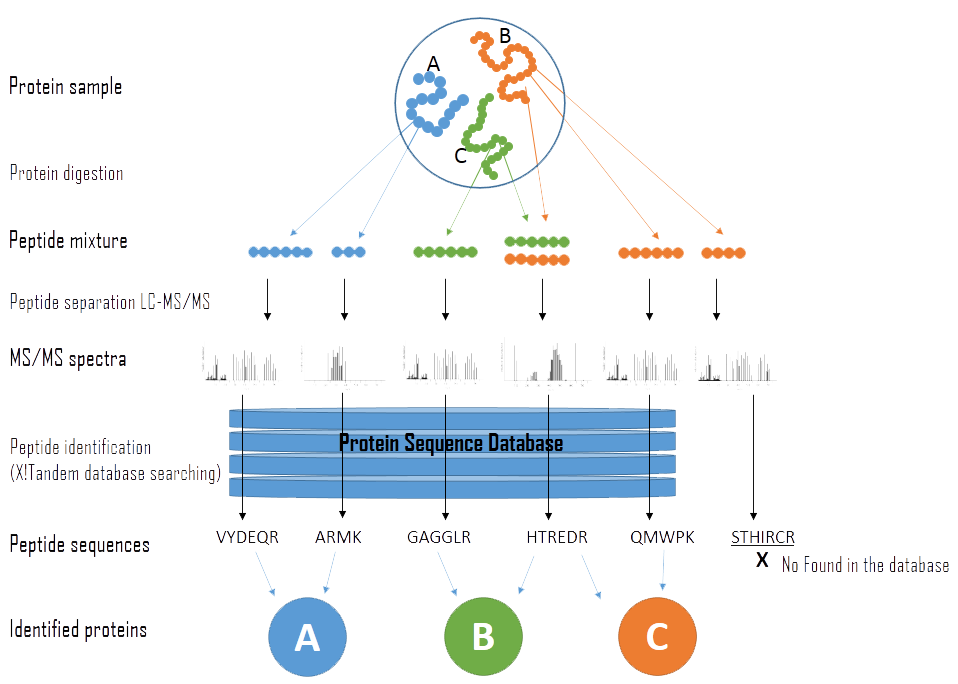
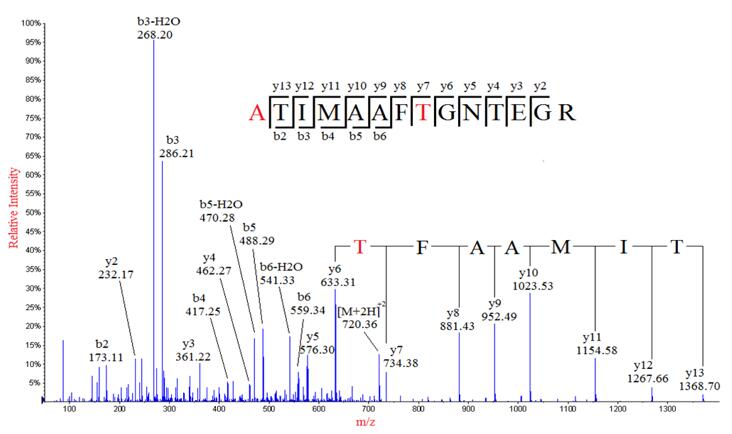
Data acquisition and database search. For a given experimental MS/MS... | Download Scientific Diagram

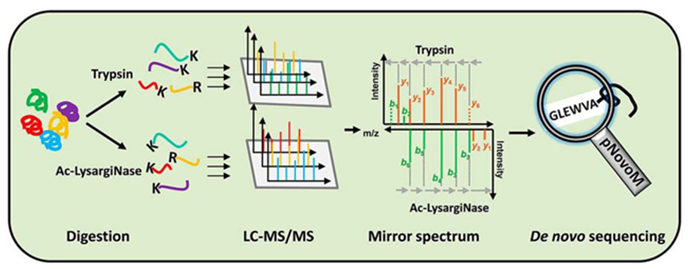
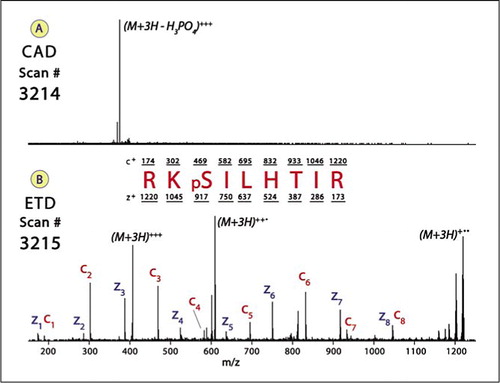
Figure 3 from Ultrahigh-resolution Fourier transform ion cyclotron resonance mass spectrometry and tandem mass spectrometry for peptide de novo amino acid sequencing for a seven-protein mixture by paired single-residue transposed Lys-N and