Dual indexed design of in-Drop single-cell RNA-seq libraries improves sequencing quality and throughput | bioRxiv

An optimized, amplicon-based approach for sequencing of SARS-CoV-2 from patient samples using COVIDSeq assay on Illumina MiSeq sequencing platforms - ScienceDirect
Large Scale Loss of Data in Low-Diversity Illumina Sequencing Libraries Can Be Recovered by Deferred Cluster Calling | PLOS ONE

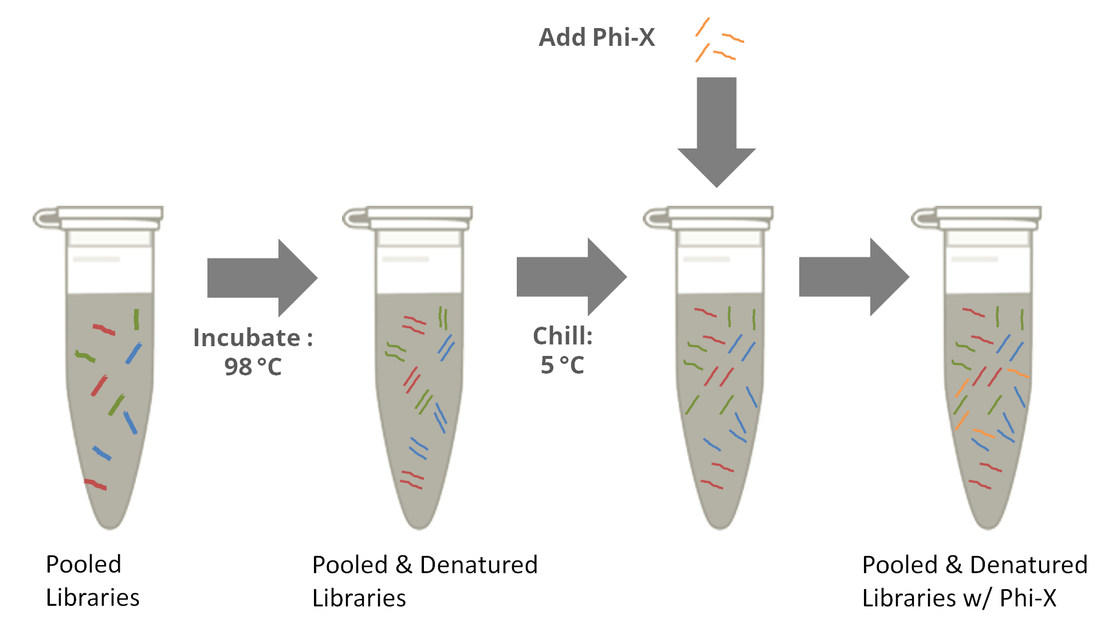
How much PhiX spike in is recommended when sequencing low diversity libraries on Illumina platforms? - Illumina Knowledge
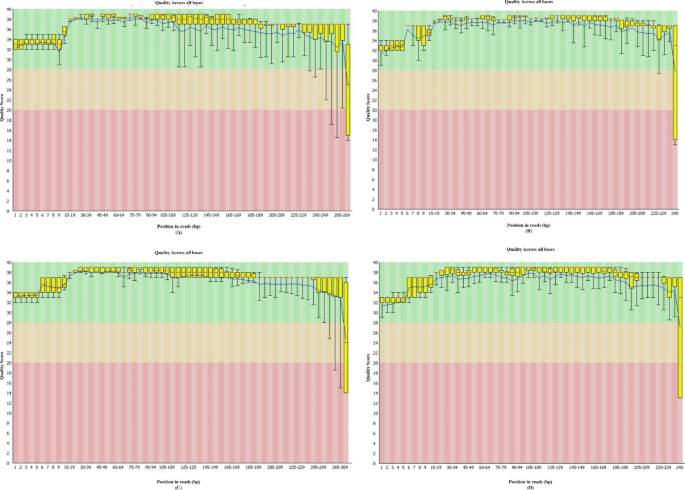
High-quality single amplicon sequencing method for illumina MiSeq platform using pool of 'N' (0–10) spacer-linked target specific primers without PhiX spike-in | BMC Genomics | Full Text

High quality single amplicon sequencing method for illumina platforms using 'N' (0-10) spacer primer pool without PhiX spik-in | bioRxiv

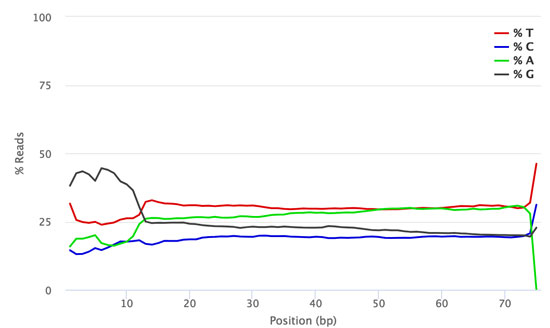

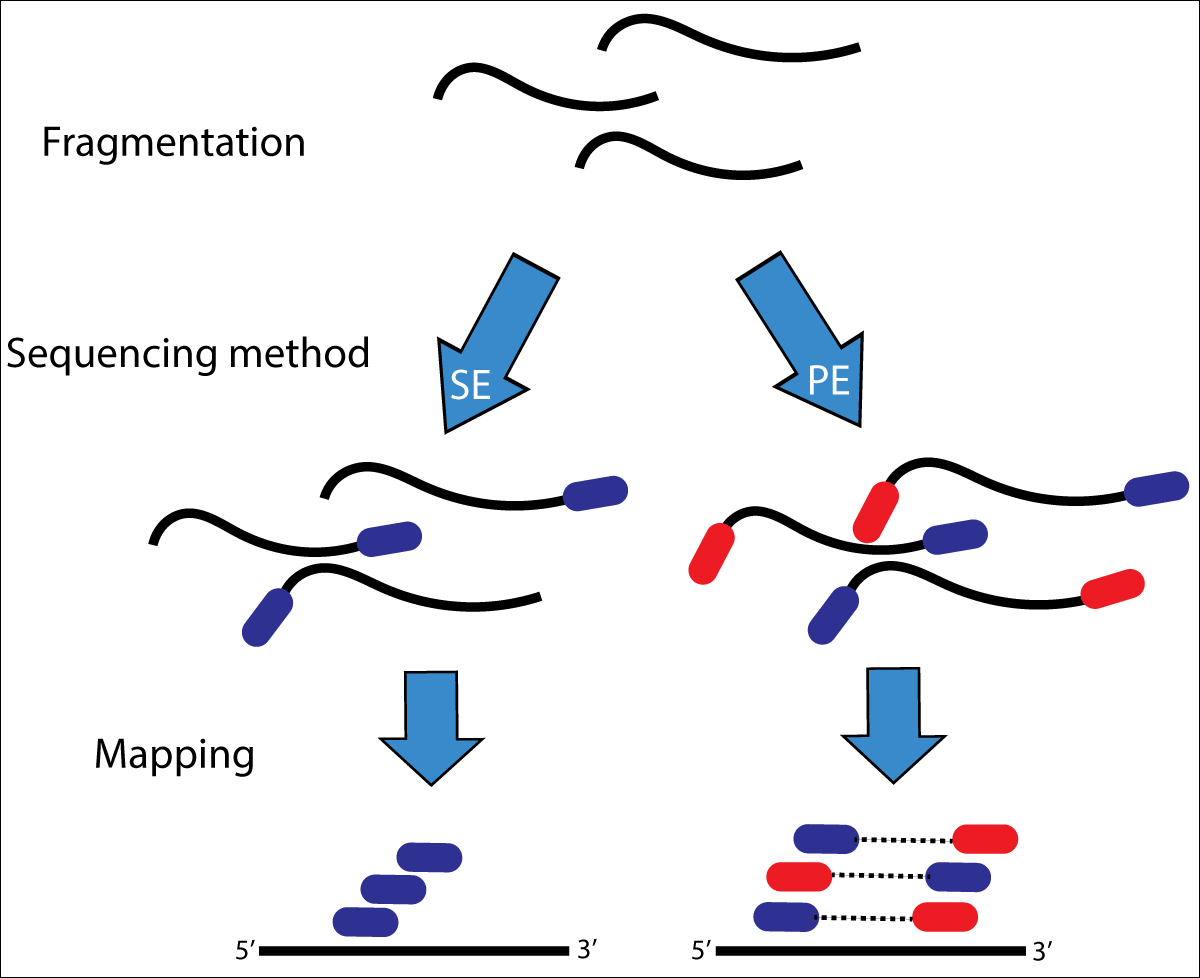
![Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ] Adapterama II: universal amplicon sequencing on Illumina platforms (TaggiMatrix) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7786/1/fig-1-full.png)