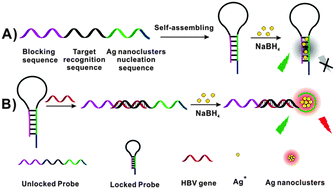
Hairpin DNA probes based on target-induced in situ generation of luminescent silver nanoclusters - Chemical Communications (RSC Publishing)
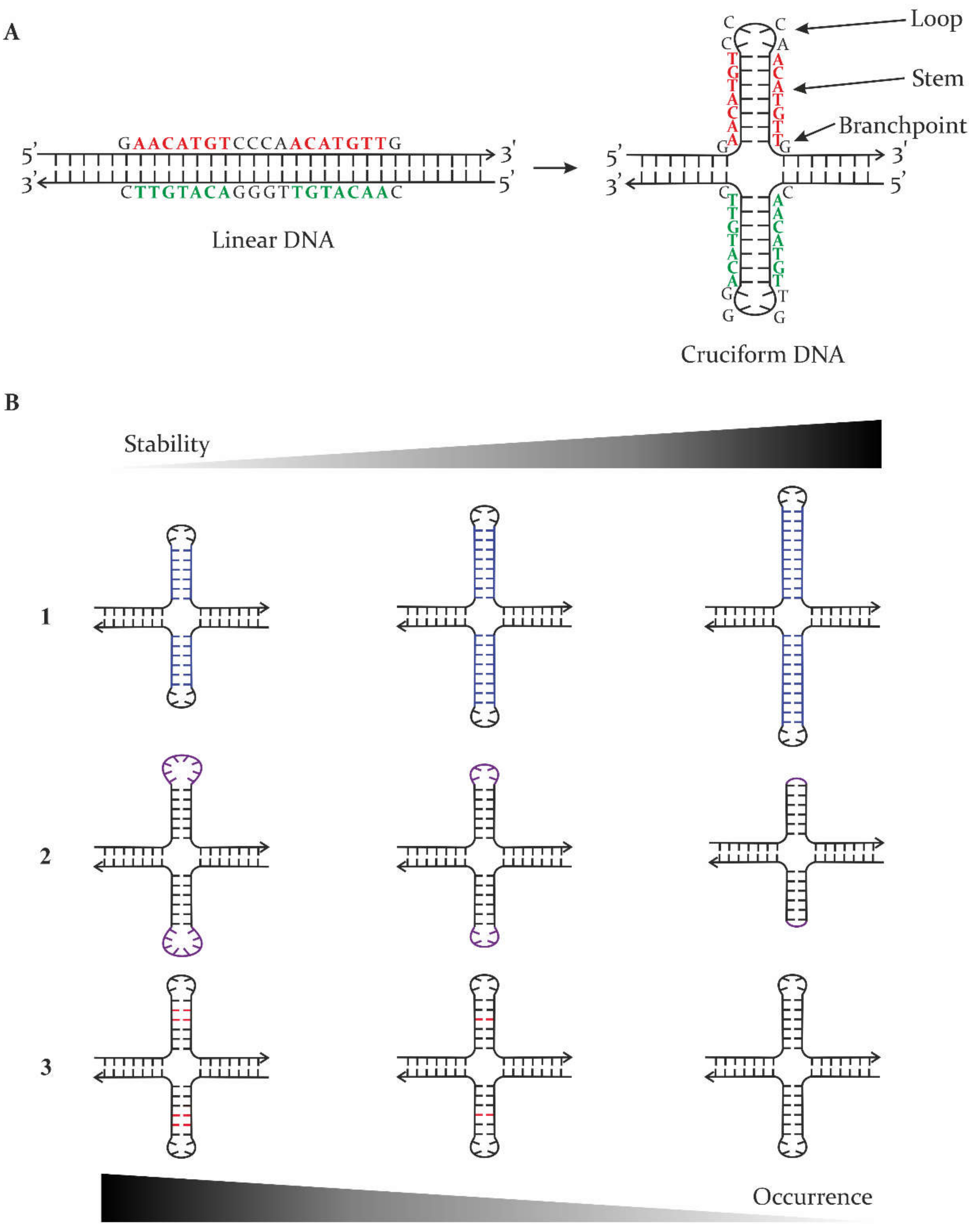
IJMS | Free Full-Text | Interaction of Proteins with Inverted Repeats and Cruciform Structures in Nucleic Acids

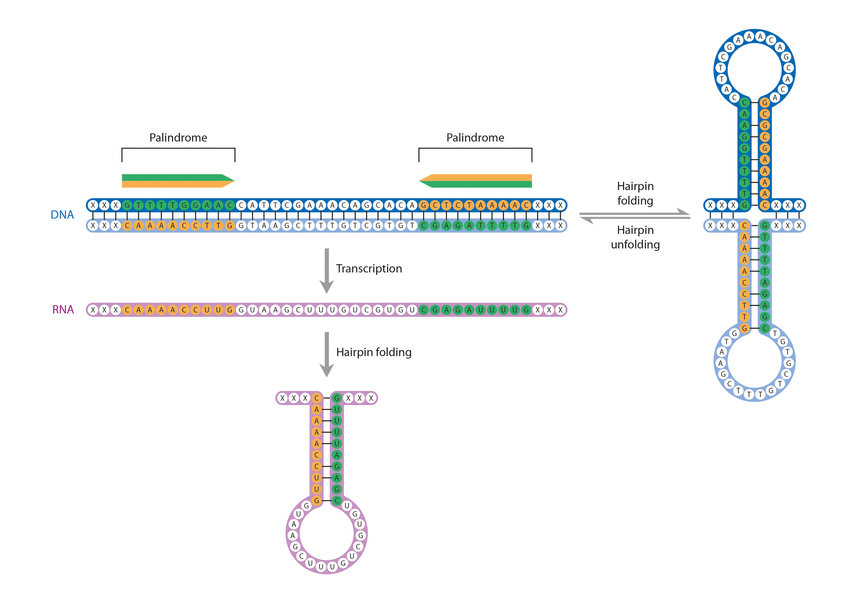
Figure 4 from Transcription termination : Nucleotide sequence at 3 ' end of tryptophan operon-in-Eseherichia coli ( DNA sequence / dyad symmetry / RNA hairpin / rho factor ) | Semantic Scholar

The Size of the Internal Loop in DNA Hairpins Influences Their Targeting with Partially Complementary Strands | The Journal of Physical Chemistry B
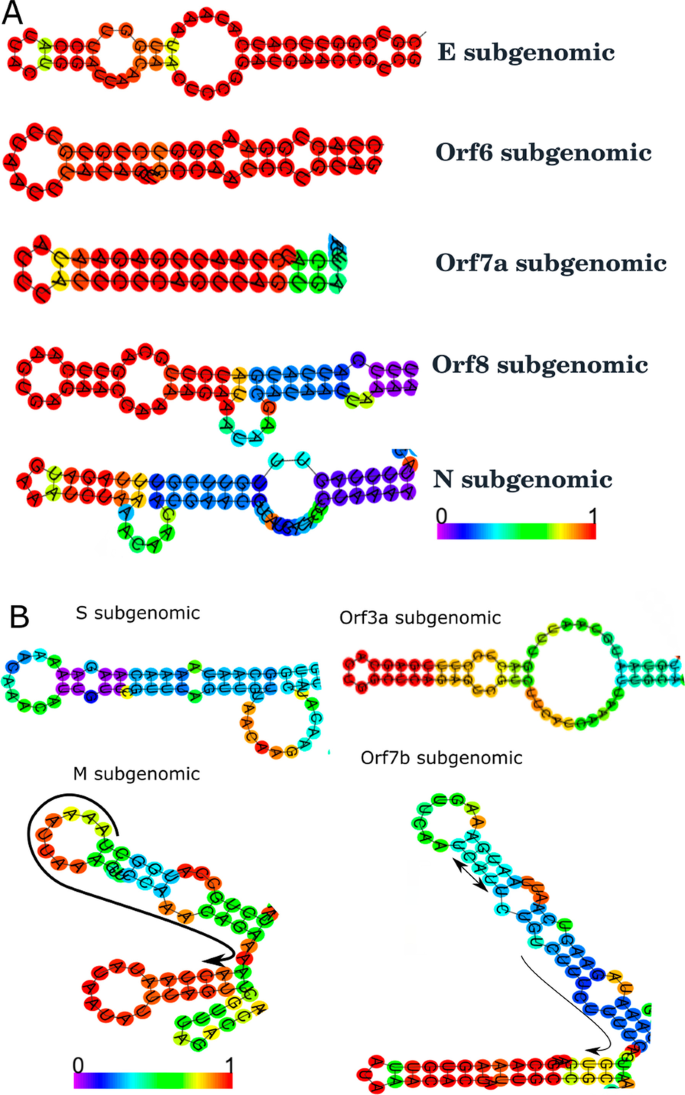
Identification of consensus hairpin loop structure among the negative sense subgenomic RNAs of SARS-CoV-2 | Bulletin of the National Research Centre | Full Text

Folded DNA in Action: Hairpin Formation and Biological Functions in Prokaryotes | Microbiology and Molecular Biology Reviews

Hairpin structures with conserved sequence motifs determine the 3′ ends of non-polyadenylated invertebrate iridovirus transcripts - ScienceDirect
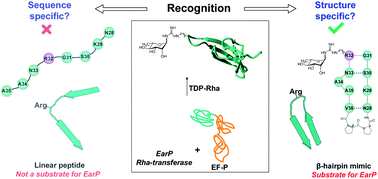
A β-hairpin epitope as novel structural requirement for protein arginine rhamnosylation - Chemical Science (RSC Publishing)

Protein Backbone Alteration in Non‐Hairpin β‐Turns: Impacts on Tertiary Folded Structure and Folded Stability - Harmon - ChemBioChem - Wiley Online Library

Core Hairpin Structure of SpCas9 sgRNA Functions in a Sequence- and Spatial Conformation–Dependent Manner - Mingjun Jiang, Yanzhen Ye, Juan Li, 2021

Folded DNA in Action: Hairpin Formation and Biological Functions in Prokaryotes | Microbiology and Molecular Biology Reviews
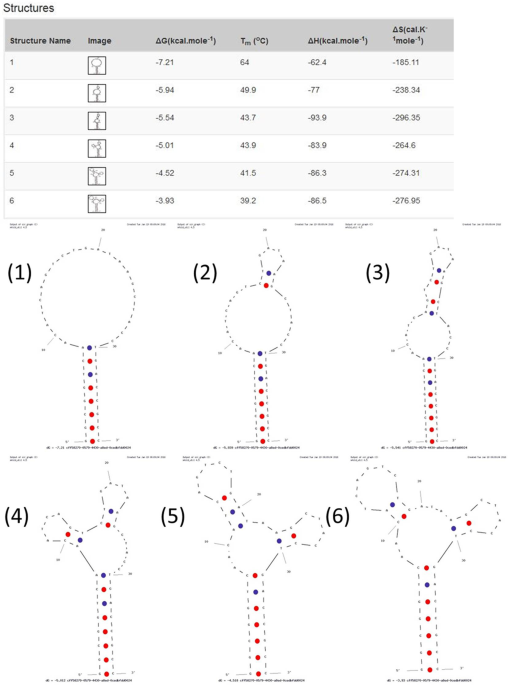
Evaluation of Rationally Designed Label-free Stem-loop DNA Probe Opening in the Presence of miR-21 by Circular Dichroism and Fluorescence Techniques | Scientific Reports