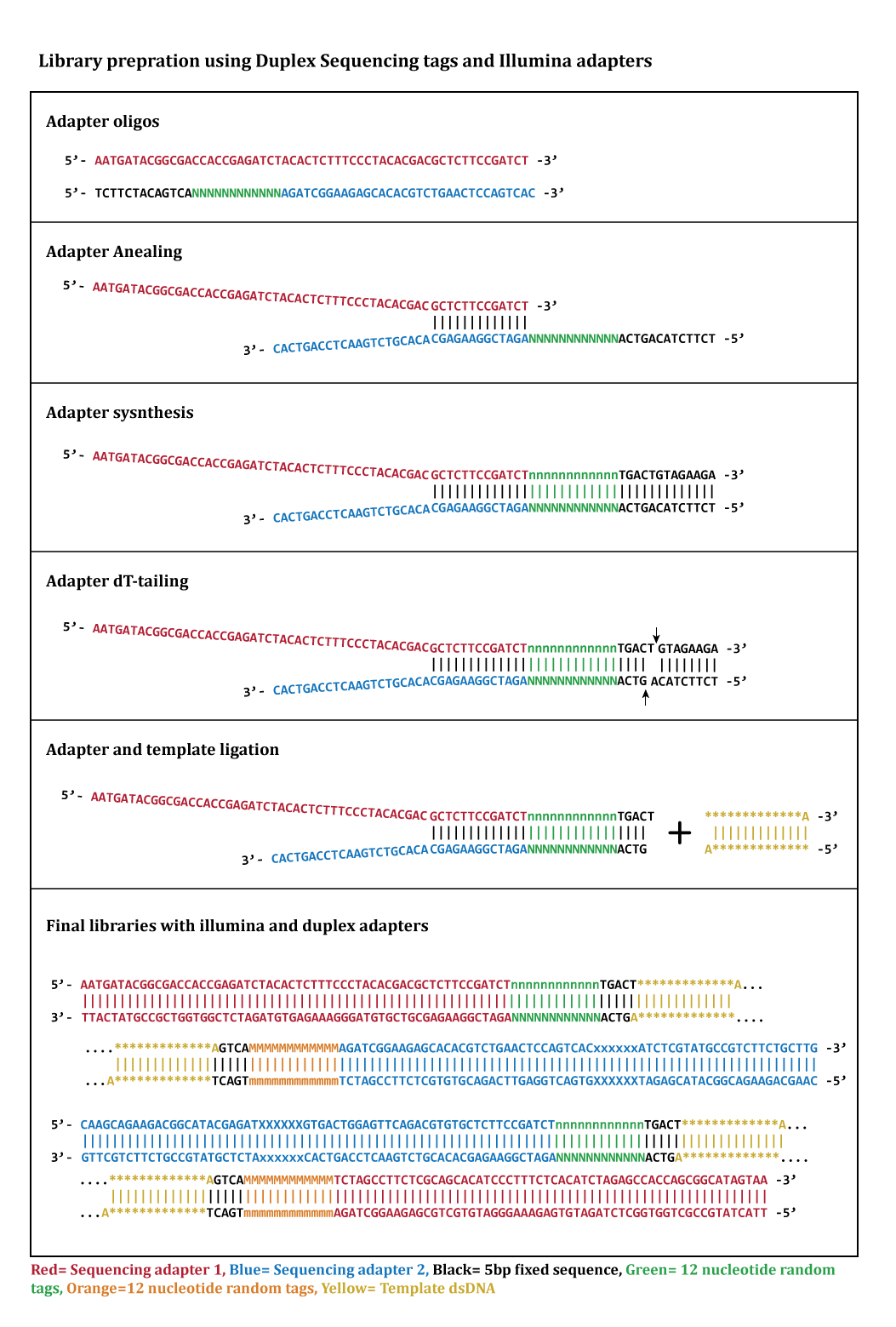
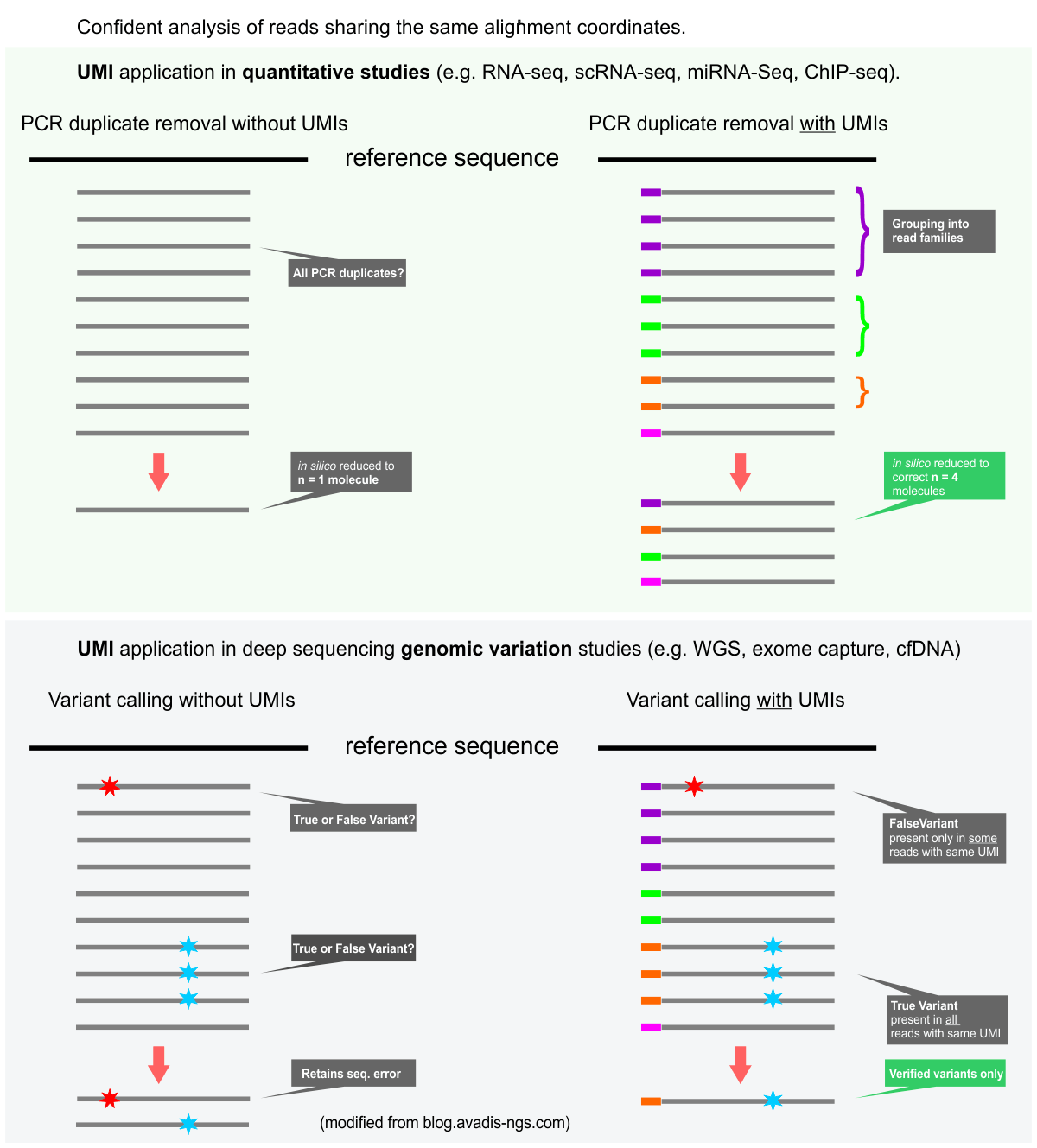
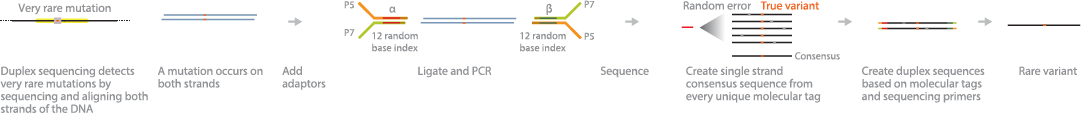Duplex Proximity Sequencing (Pro-Seq): A method to improve DNA sequencing accuracy without the cost of molecular barcoding redundancy | PLOS ONE

TwinStrand's Duplex Sequencing Goes Deep: When it comes to detection of low frequency variants, Jesse Salk's technology is one in (ten) million: GEN Edge: Vol 2, No 1

SinoDuplex: An Improved Duplex Sequencing Approach to Detect Low-frequency Variants in Plasma cfDNA Samples | Semantic Scholar

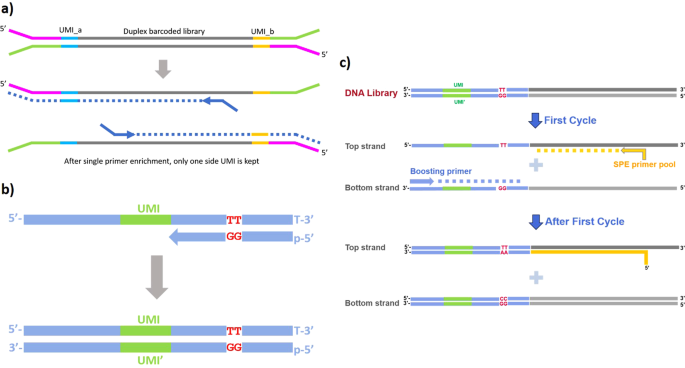
Design of single-end duplex-UMI adapter. (a) Schematics showing how... | Download Scientific Diagram

Estimating somatic mutation rates by Duplex Sequencing in non-model organisms: Daphnia magna as a case study | bioRxiv
Research Sequencing Service: Supported Analysis Pipelines | Research | Dept. of Laboratory Medicine & Pathology | UW Medicine

Inigo Martincorena on Twitter: "Over 4 years of iterative refinement, Rob Osborne and @AbascalFed identified sources of error in duplex sequencing: end repair, nick extension, mapping errors, DNA contamination… devising solutions to
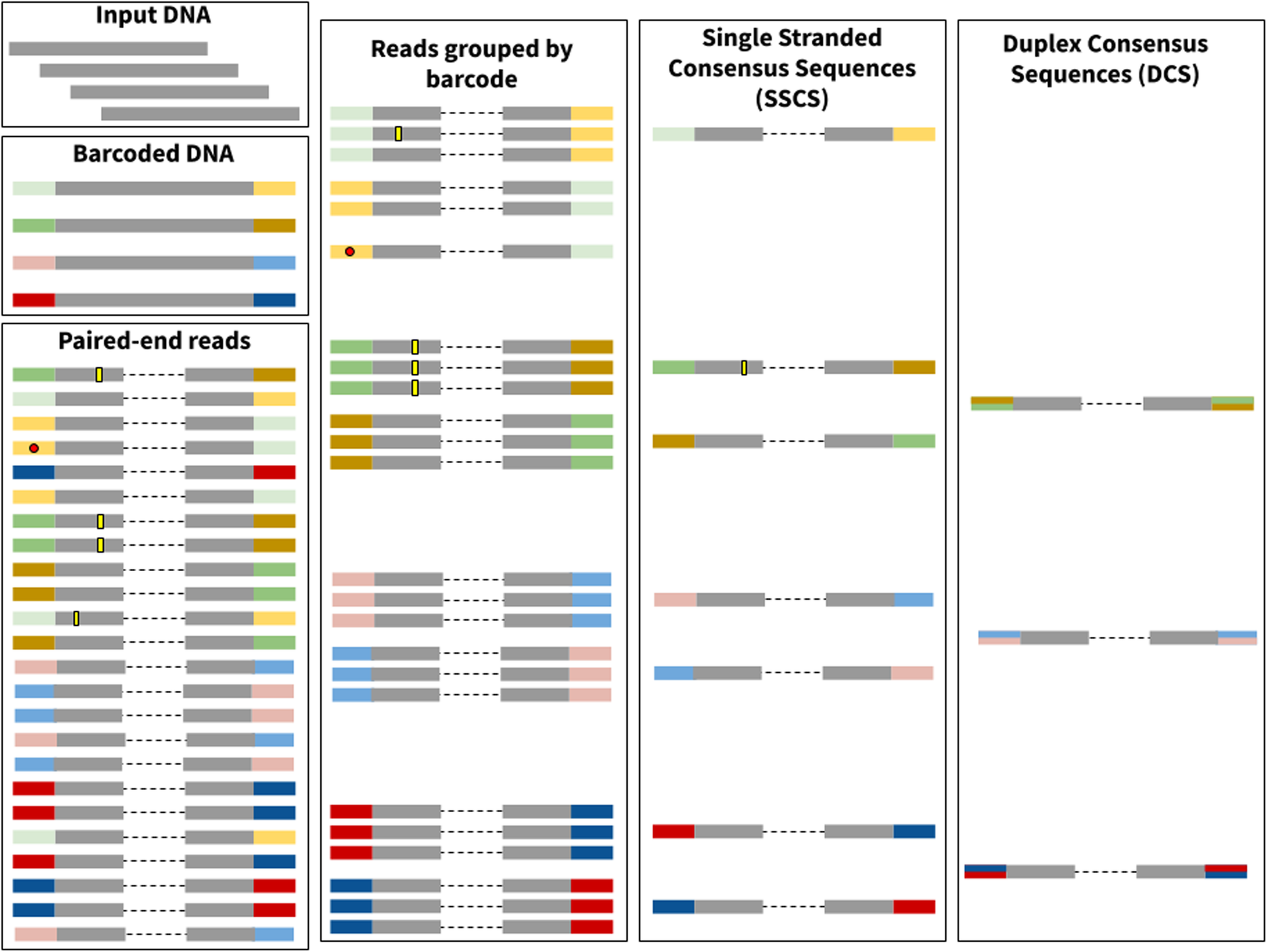

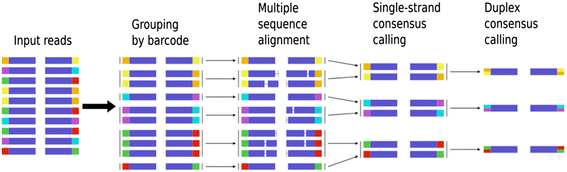
Family reunion via error correction: an efficient analysis of duplex sequencing data | BMC Bioinformatics | Full Text