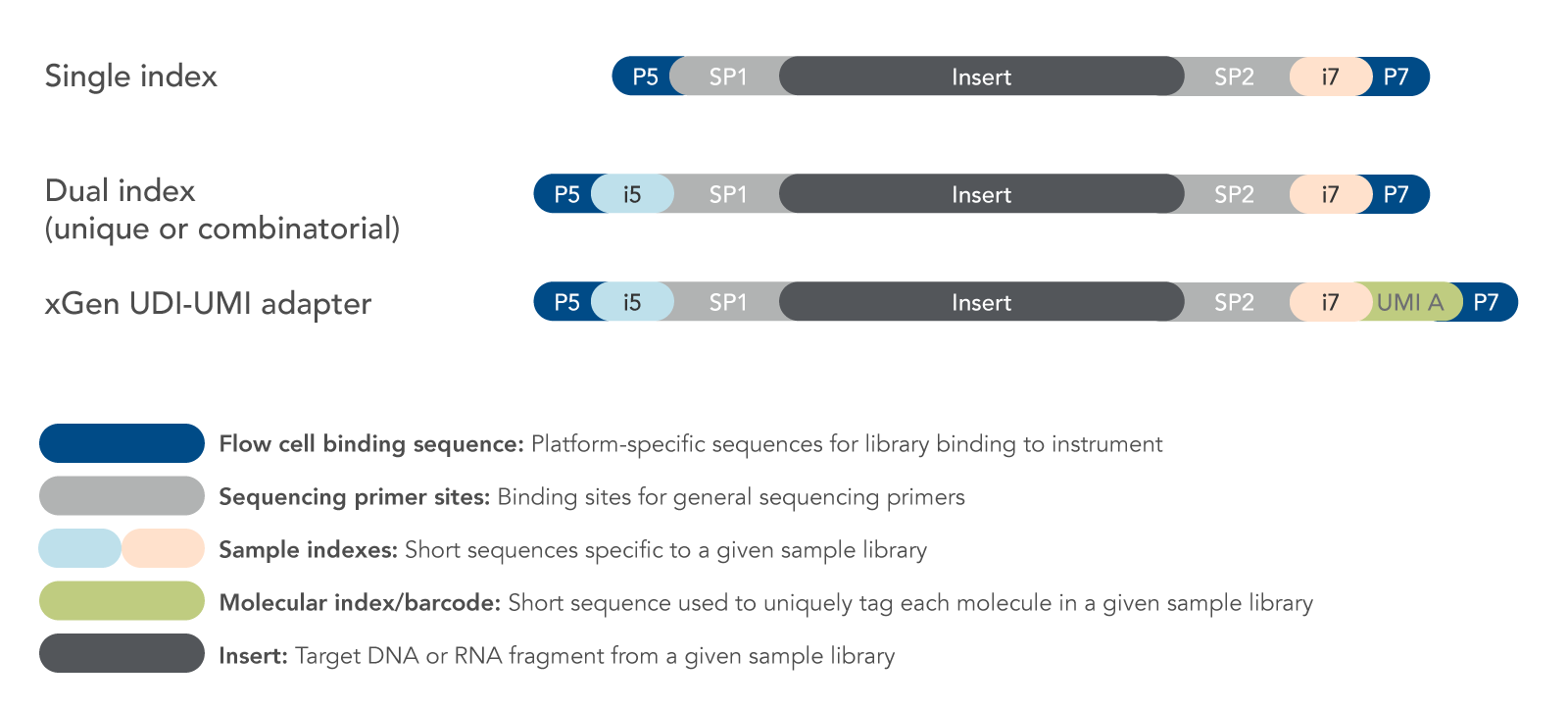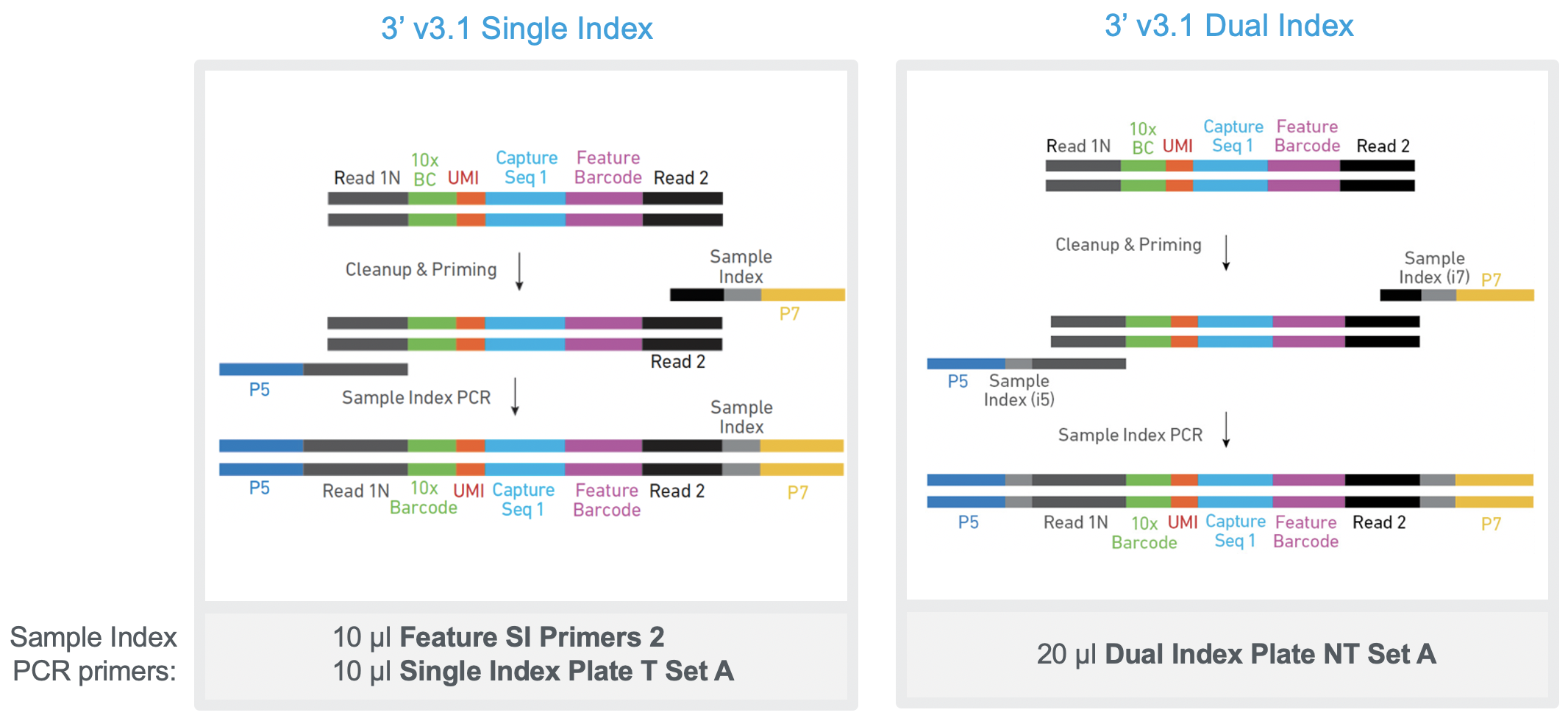
What are the protocol differences between Single Cell 3' v3.1 Single Index and 3' v3.1 Dual Index? – 10X Genomics

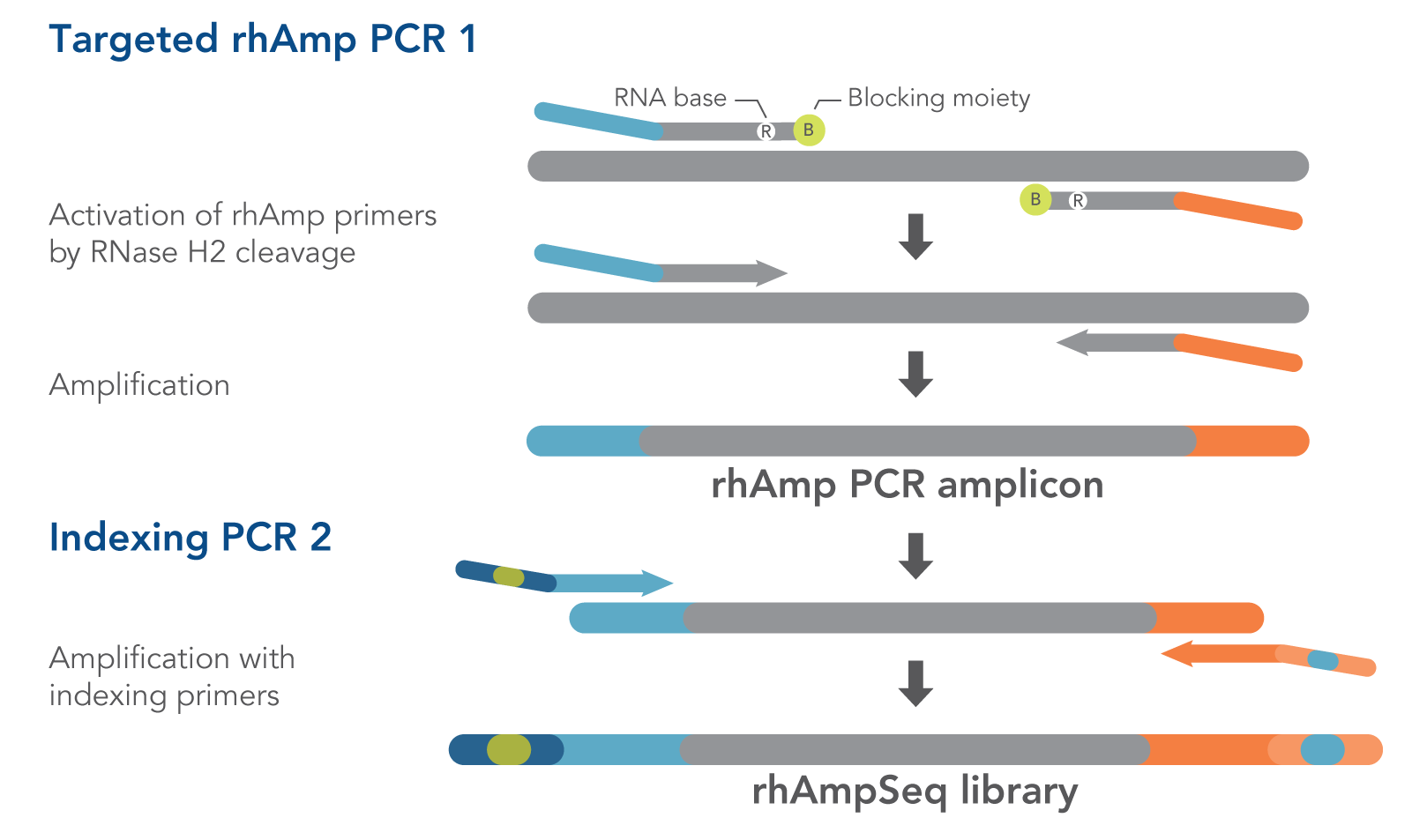
Ultrahigh-Throughput Multiplexing and Sequencing of >500-Base-Pair Amplicon Regions on the Illumina HiSeq 2500 Platform | mSystems
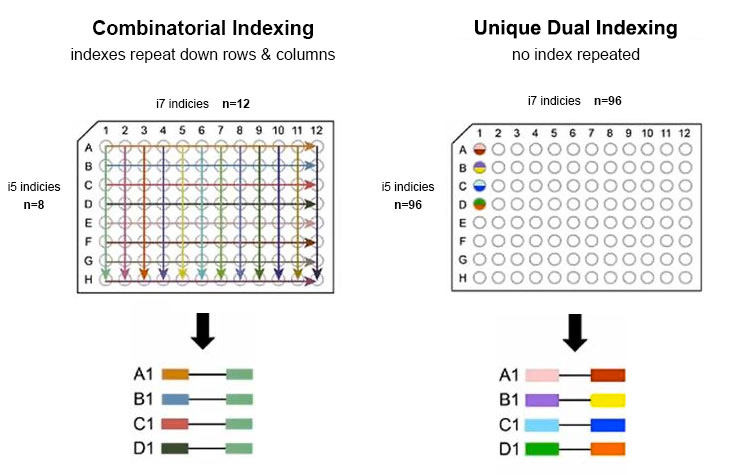
![Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ] Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7755/1/fig-1-full.png)
Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]
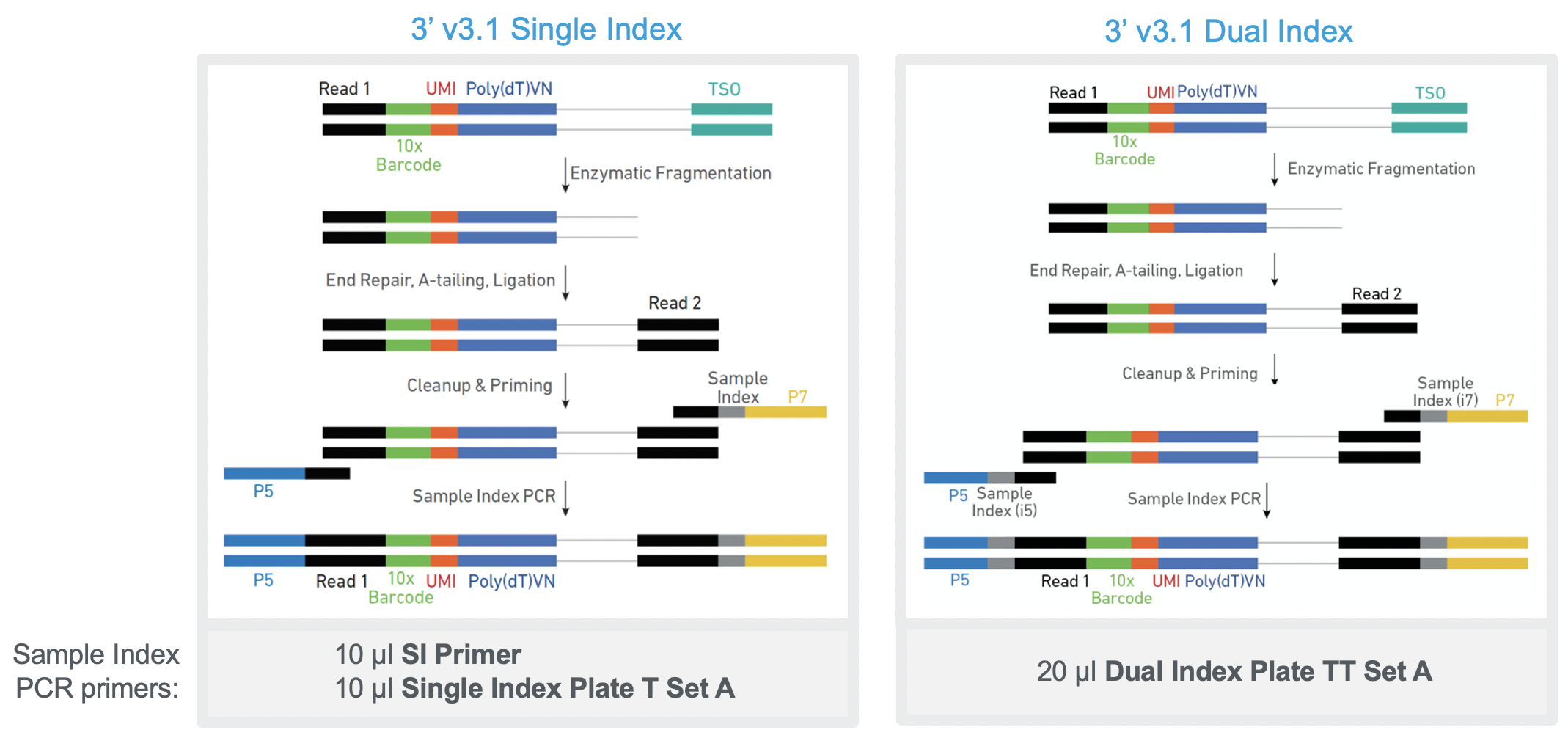
What are the protocol differences between Single Cell 3' v3.1 Single Index and 3' v3.1 Dual Index? – 10X Genomics

Unique dual indexing PCR reduces chimeric contamination and improves mutation detection in cell-free DNA of pregnant women - ScienceDirect
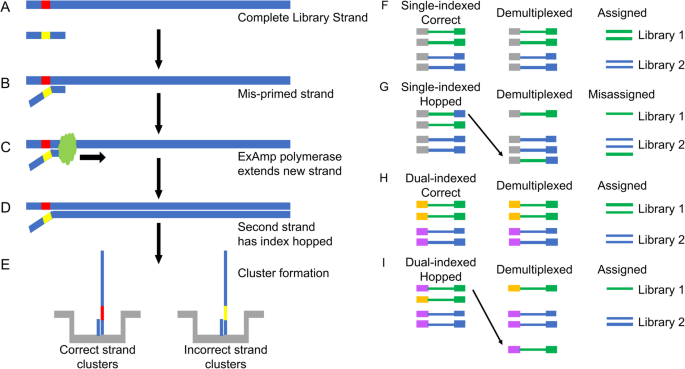
Dual indexed library design enables compatibility of in-Drop single-cell RNA-sequencing with exAMP chemistry sequencing platforms | BMC Genomics | Full Text
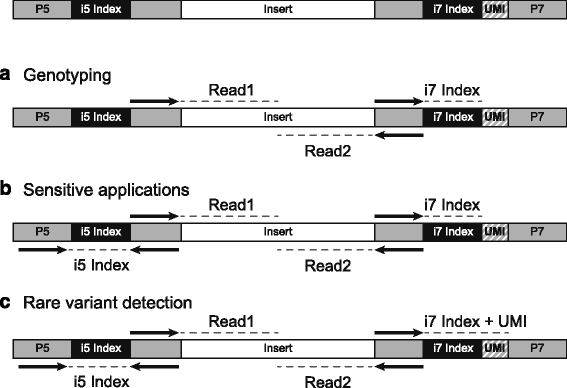
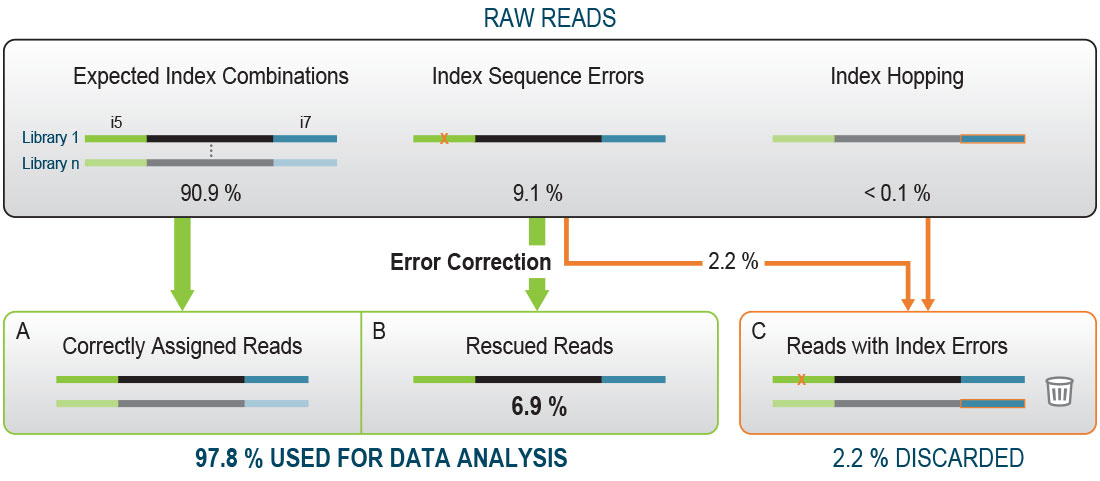
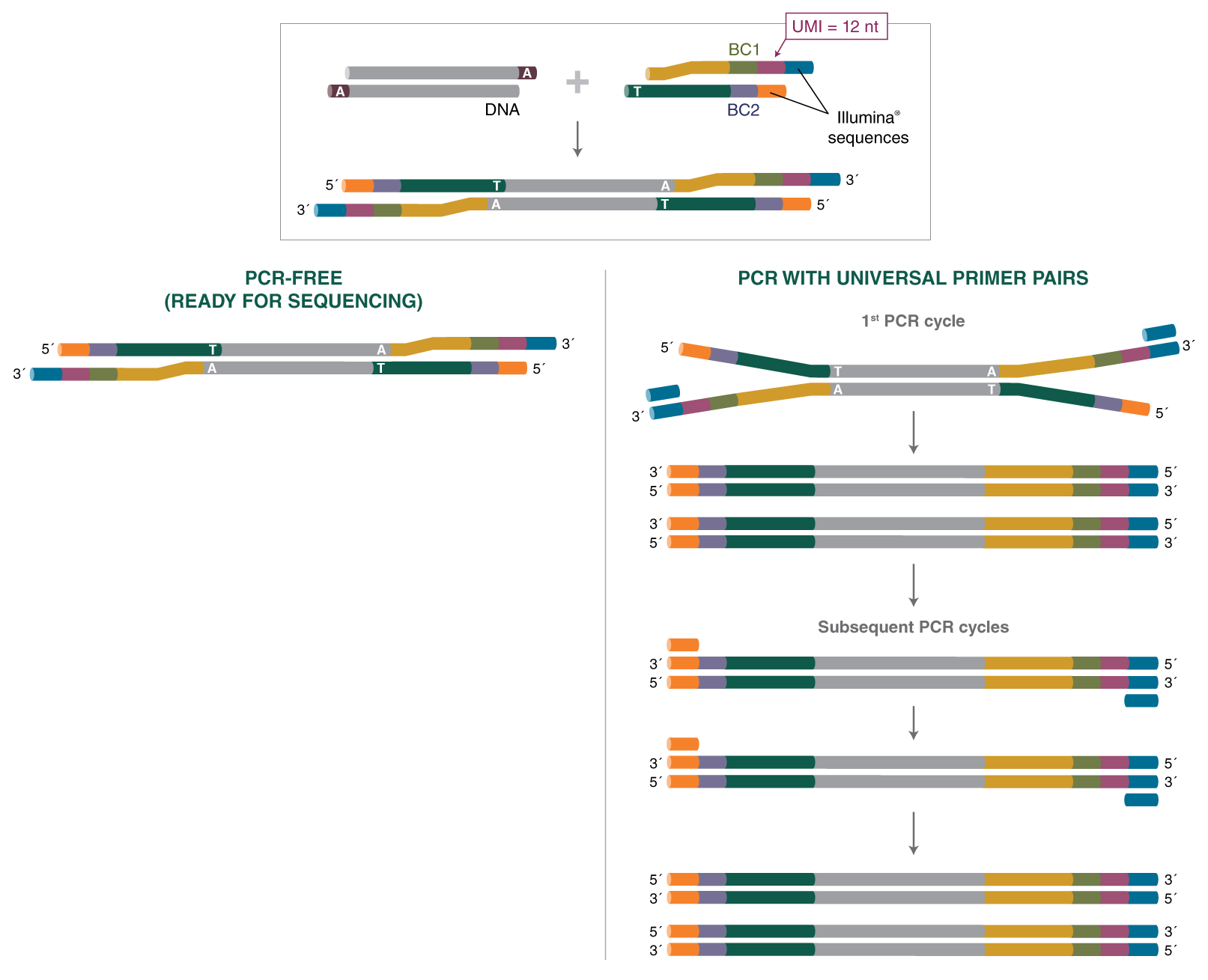
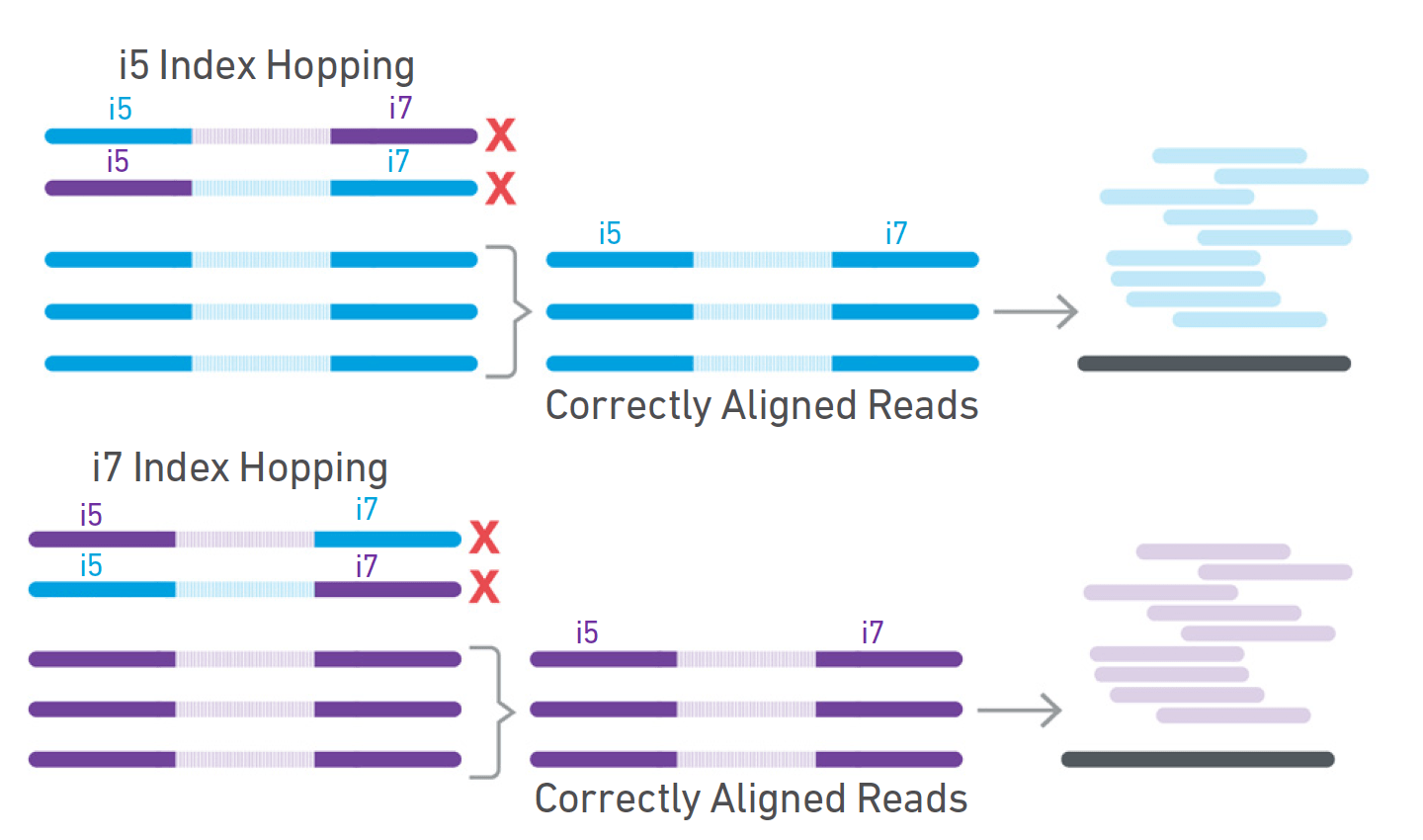
Unique, dual-indexed sequencing adapters with UMIs effectively eliminate index cross-talk and significantly improve sensitivity of massively parallel sequencing | BMC Genomics | Full Text

Dual indexed design of in-Drop single-cell RNA-seq libraries improves sequencing quality and throughput | bioRxiv

Experimental design. (a) Design of single and dual-index sequencing... | Download Scientific Diagram

Comparison of LUMI-PCR with regular Illumina dual index library prep... | Download Scientific Diagram
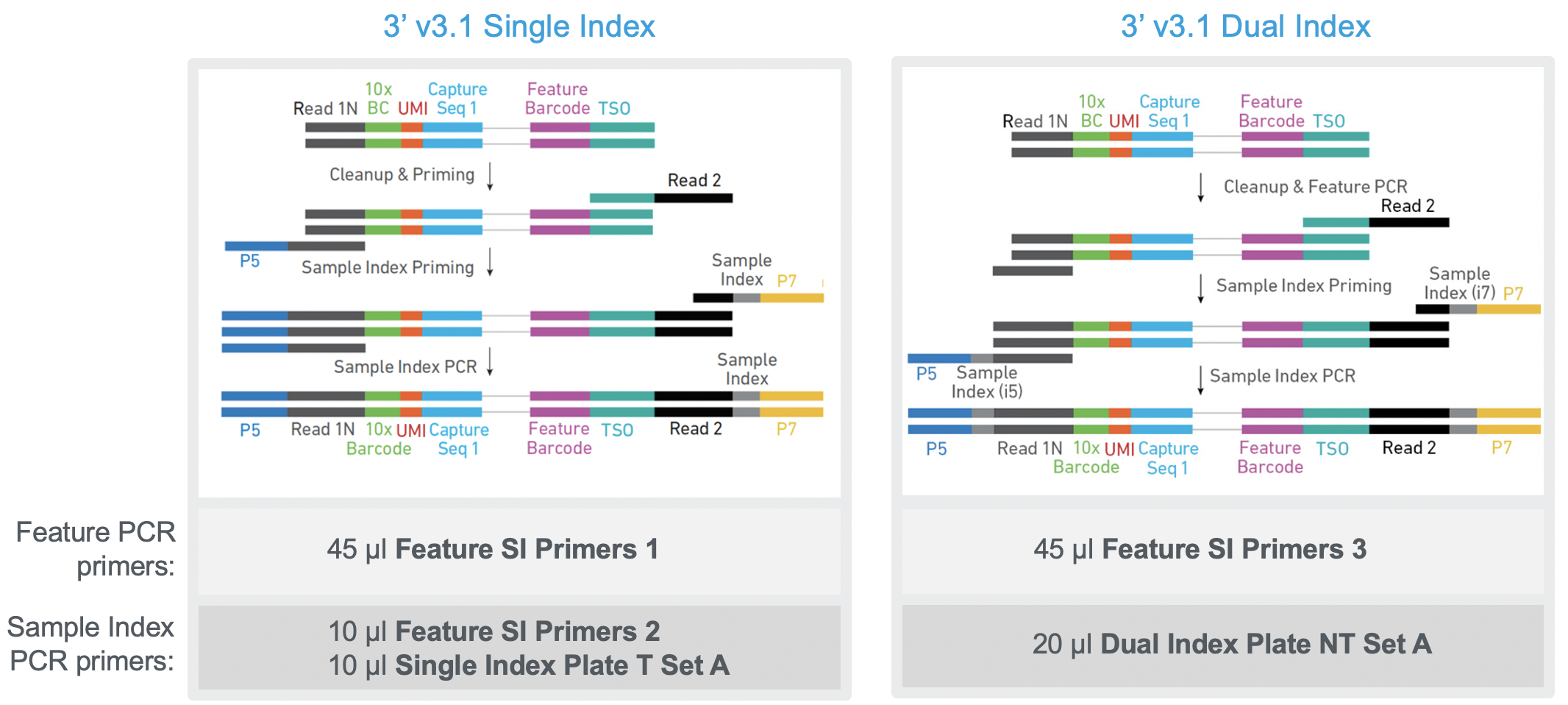
What are the protocol differences between Single Cell 3' v3.1 Single Index and 3' v3.1 Dual Index? – 10X Genomics
![Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ] Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7755/1/fig-4-2x.jpg)
Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]
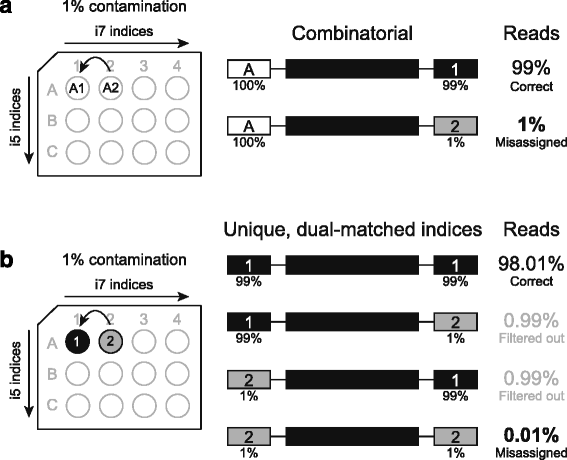
Unique, dual-indexed sequencing adapters with UMIs effectively eliminate index cross-talk and significantly improve sensitivity of massively parallel sequencing | BMC Genomics | Full Text