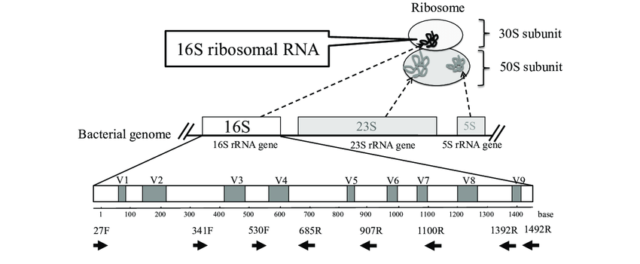
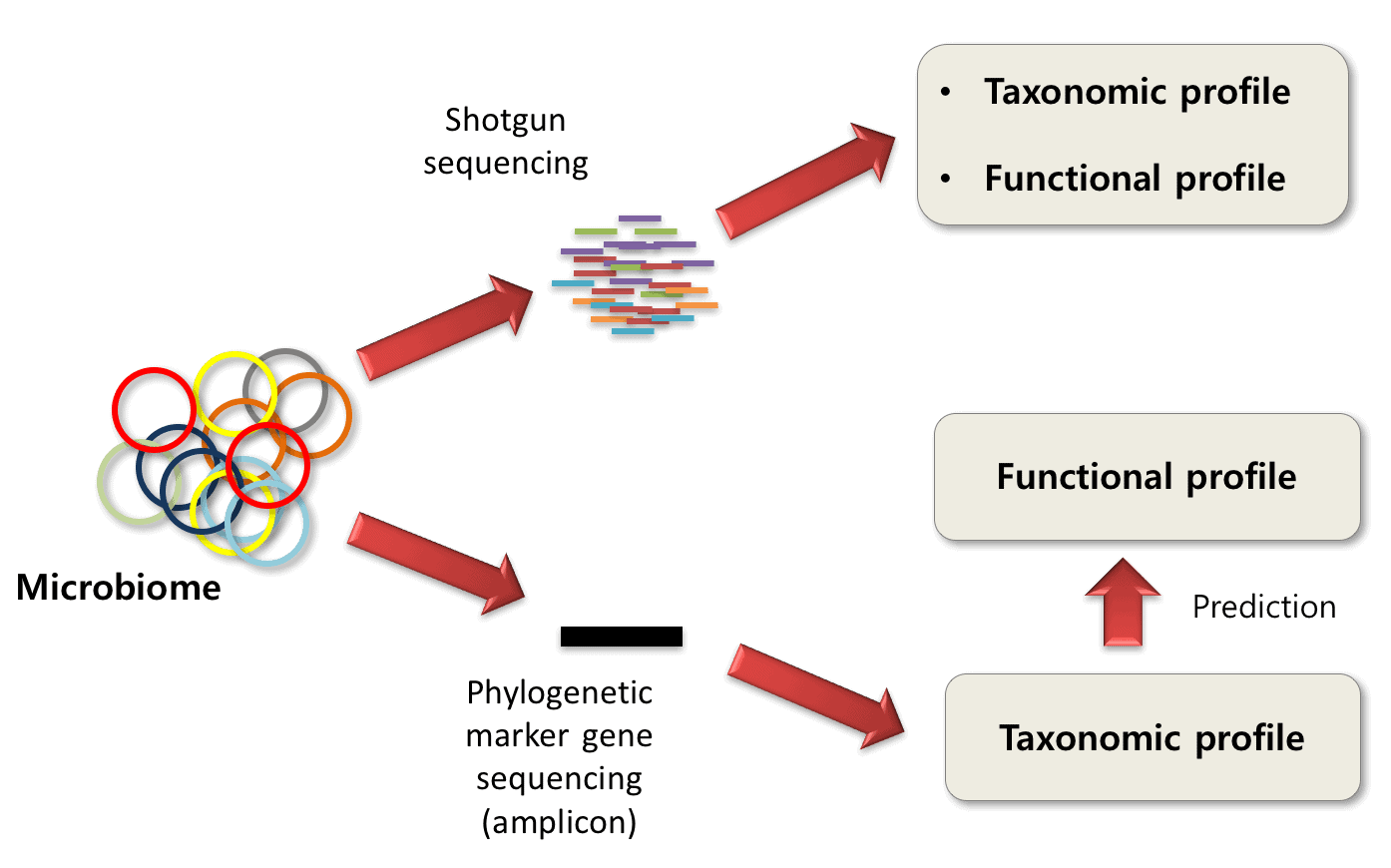
Optimization of the 16S rRNA sequencing analysis pipeline for studying in vitro communities of gut commensals - ScienceDirect
![PDF] Streamlined processing and analysis of 16S rRNA amplicon sequencing data with OCMS_16S and OCMSlooksy | Semantic Scholar PDF] Streamlined processing and analysis of 16S rRNA amplicon sequencing data with OCMS_16S and OCMSlooksy | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/07aade26e16497cbd9eb774ceb664fff4dd5aa1b/5-Figure1-1.png)
PDF] Streamlined processing and analysis of 16S rRNA amplicon sequencing data with OCMS_16S and OCMSlooksy | Semantic Scholar

EasyAmplicon: An easy‐to‐use, open‐source, reproducible, and community‐based pipeline for amplicon data analysis in microbiome research - Liu - 2023 - iMeta - Wiley Online Library

Handling of spurious sequences affects the outcome of high-throughput 16S rRNA gene amplicon profiling | ISME Communications
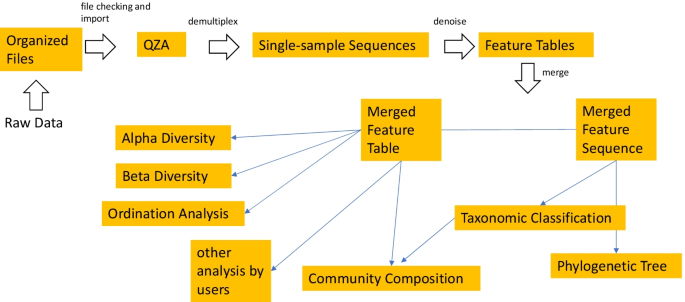
ASAP 2: a pipeline and web server to analyze marker gene amplicon sequencing data automatically and consistently | BMC Bioinformatics | Full Text

NanoRTax, a real-time pipeline for taxonomic and diversity analysis of nanopore 16S rRNA amplicon sequencing data - ScienceDirect
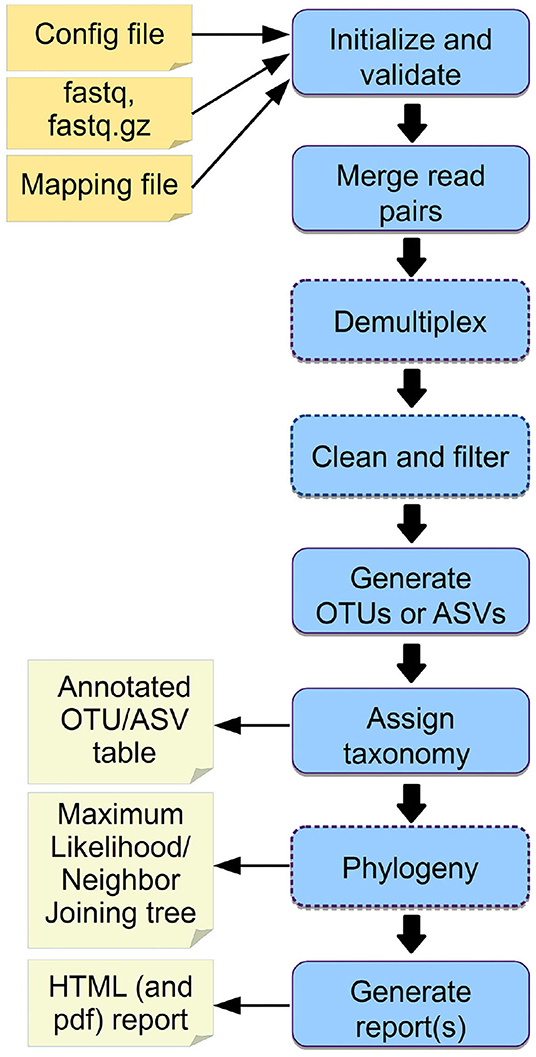
Frontiers | Cascabel: A Scalable and Versatile Amplicon Sequence Data Analysis Pipeline Delivering Reproducible and Documented Results
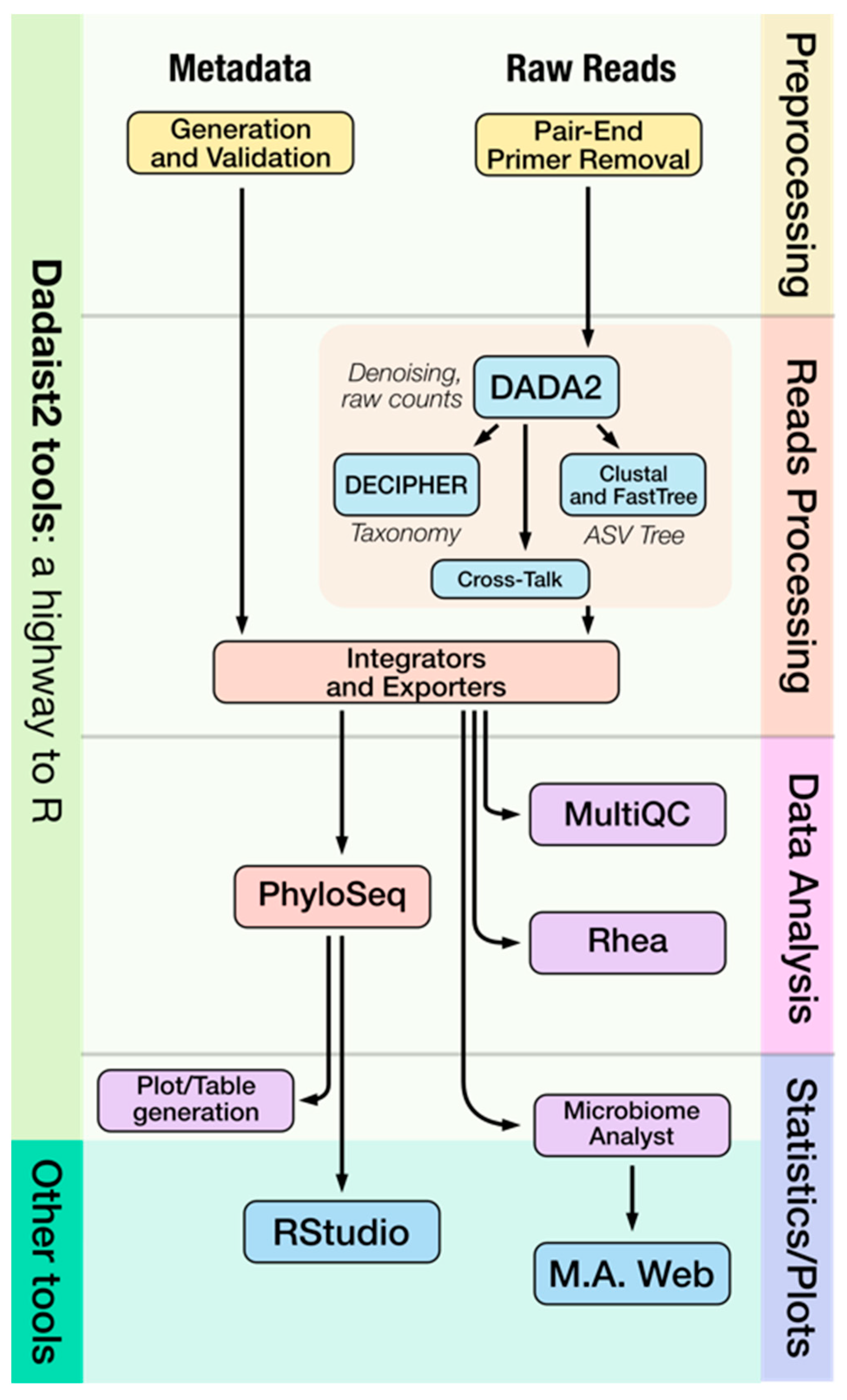
IJMS | Free Full-Text | Dadaist2: A Toolkit to Automate and Simplify Statistical Analysis and Plotting of Metabarcoding Experiments
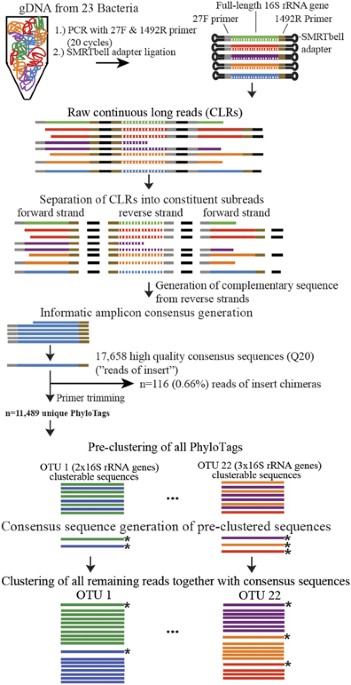
![16S-FASAS: an integrated pipeline for synthetic full-length 16S rRNA gene sequencing data analysis [PeerJ] 16S-FASAS: an integrated pipeline for synthetic full-length 16S rRNA gene sequencing data analysis [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2022/14043/1/fig-1-2x.jpg)
16S-FASAS: an integrated pipeline for synthetic full-length 16S rRNA gene sequencing data analysis [PeerJ]
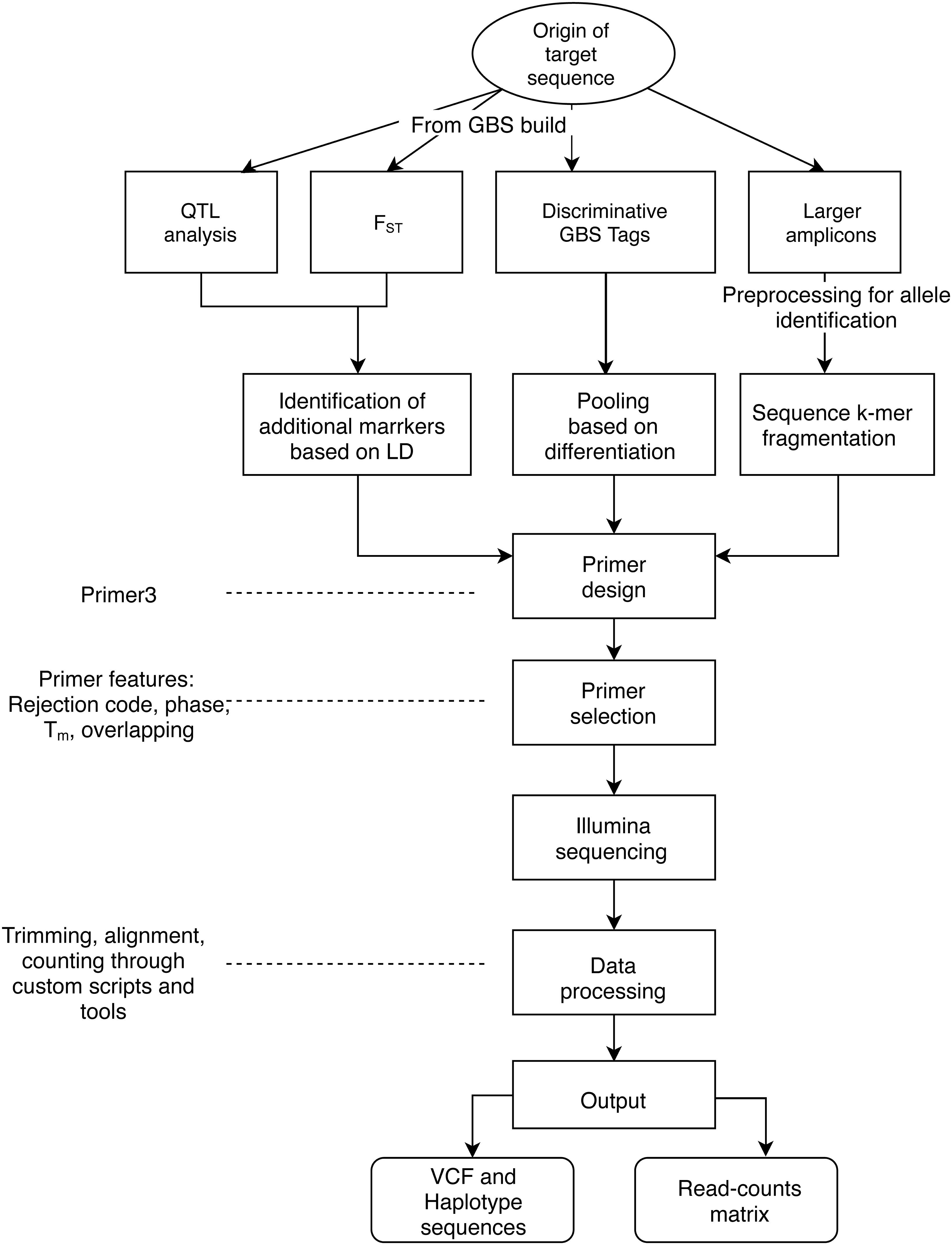
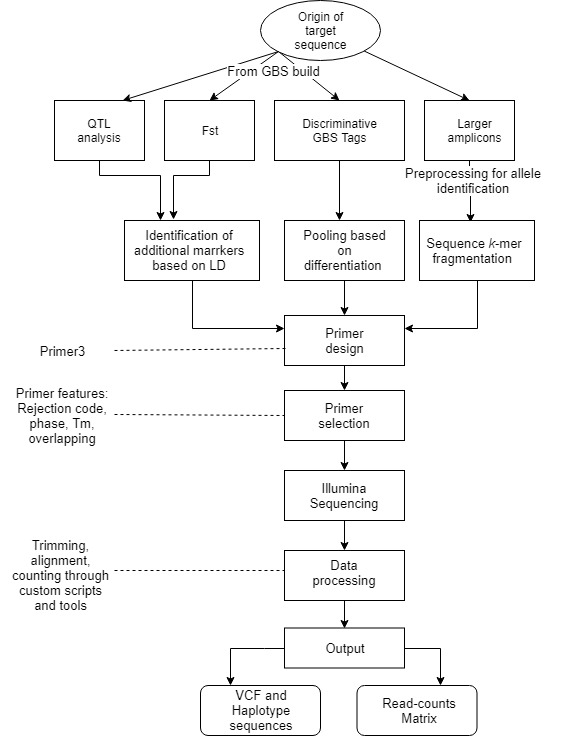
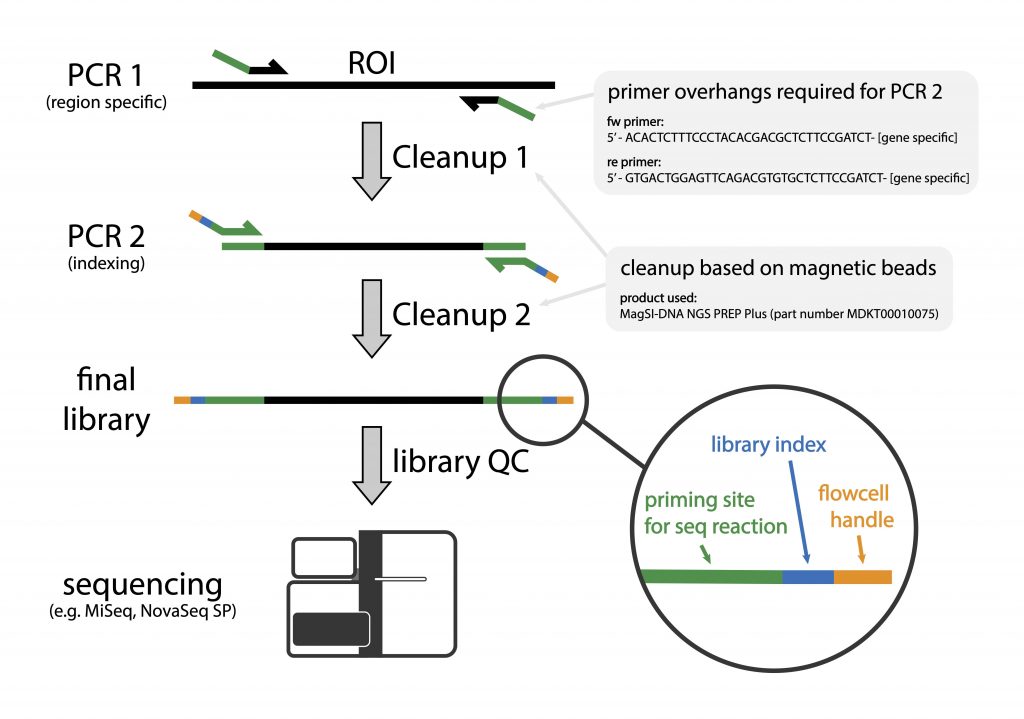
Frontiers | Computational Analysis of AmpSeq Data for Targeted, High-Throughput Genotyping of Amplicons
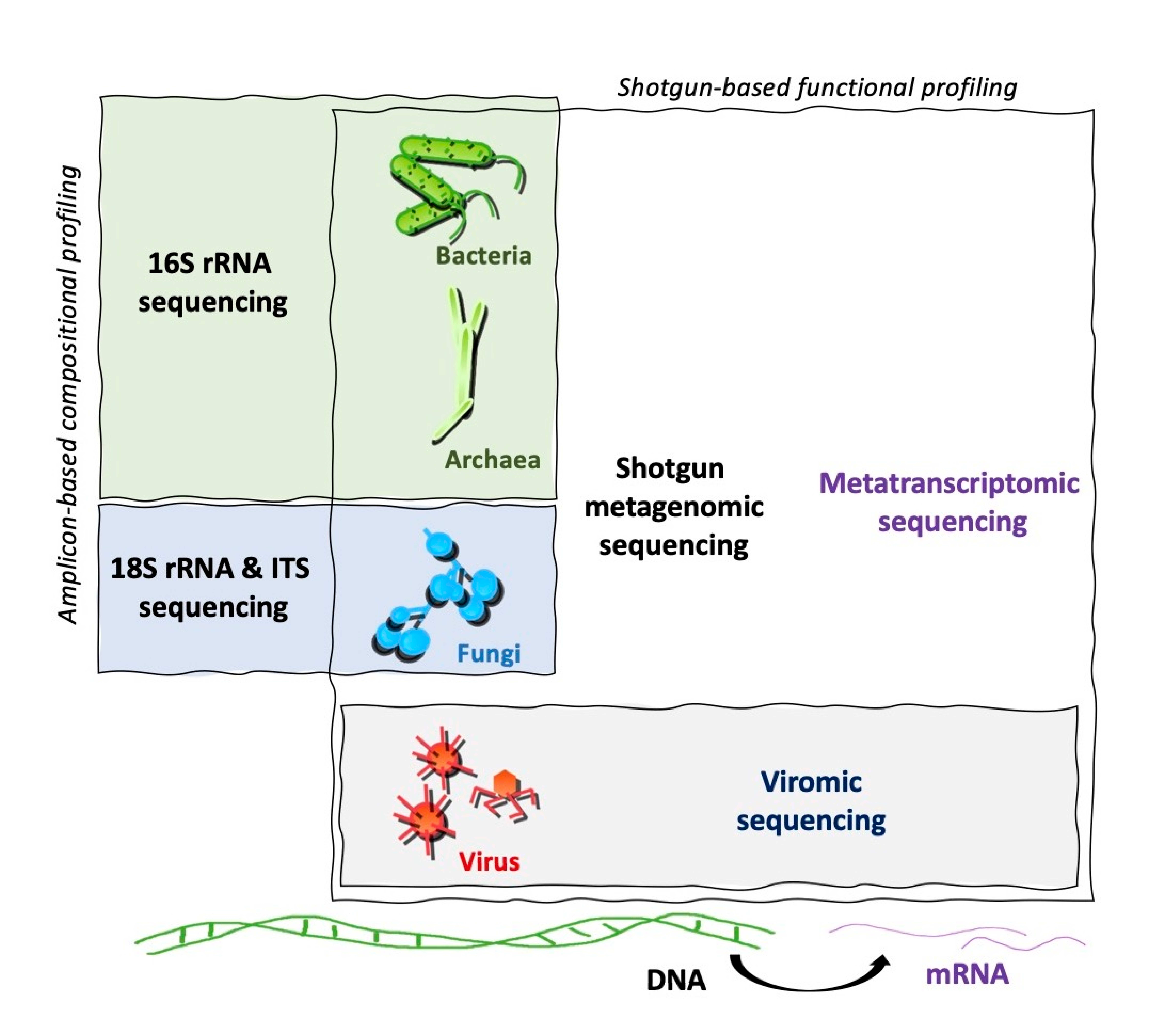
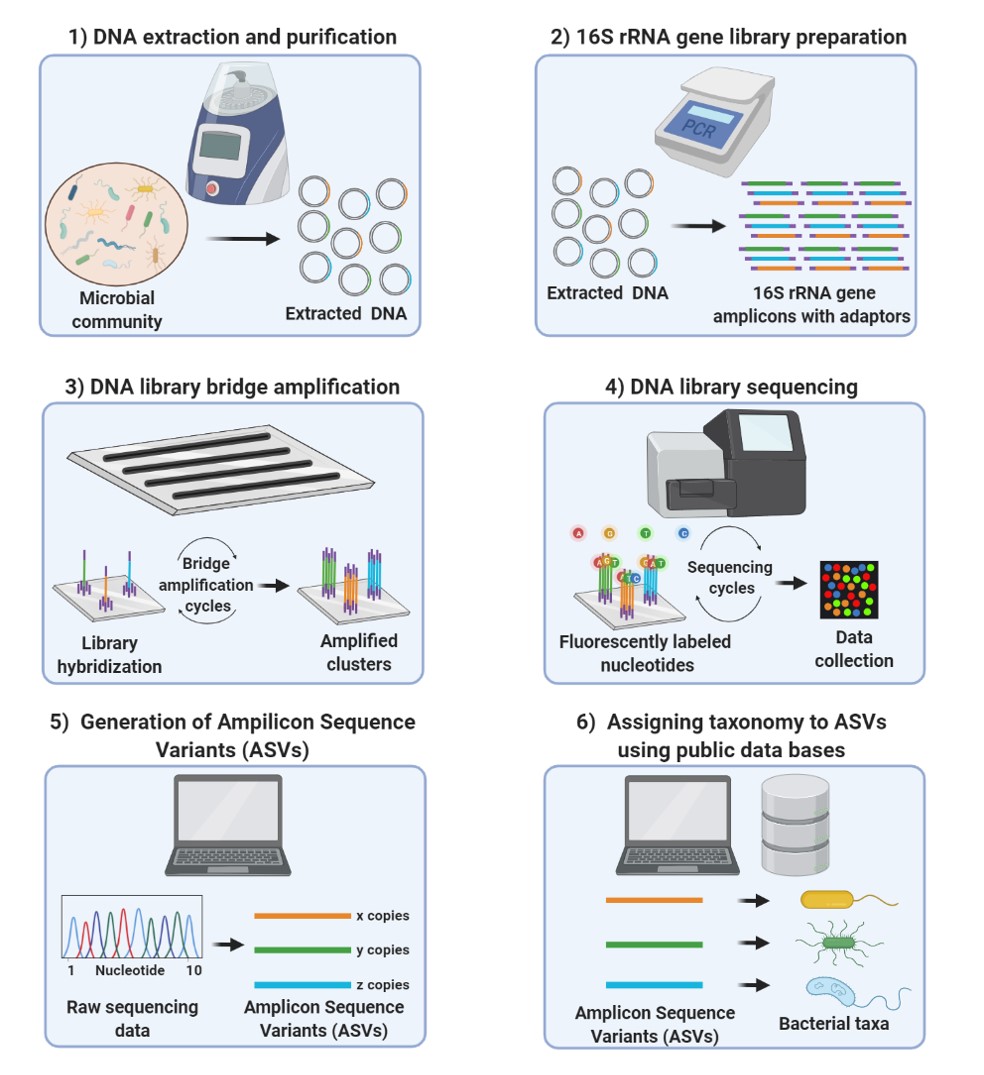
Biomolecules | Free Full-Text | An Introduction to Next Generation Sequencing Bioinformatic Analysis in Gut Microbiome Studies